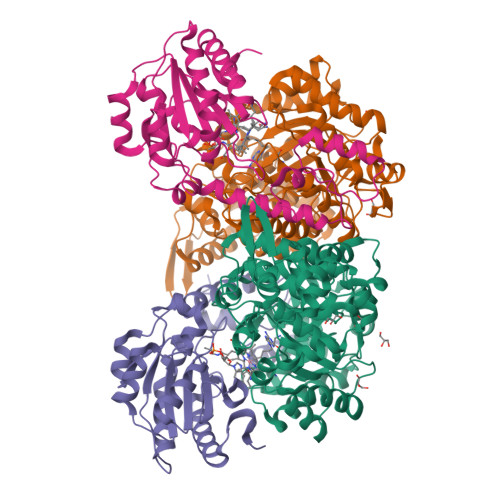
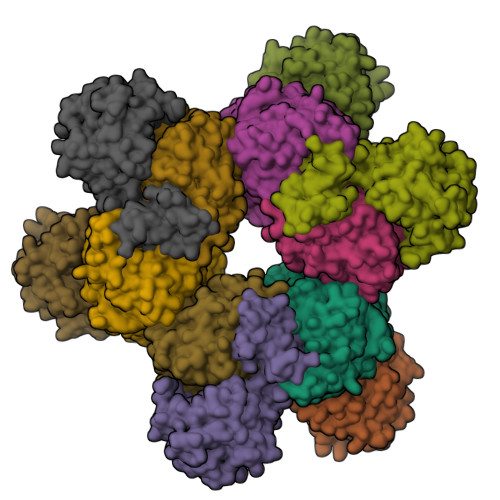
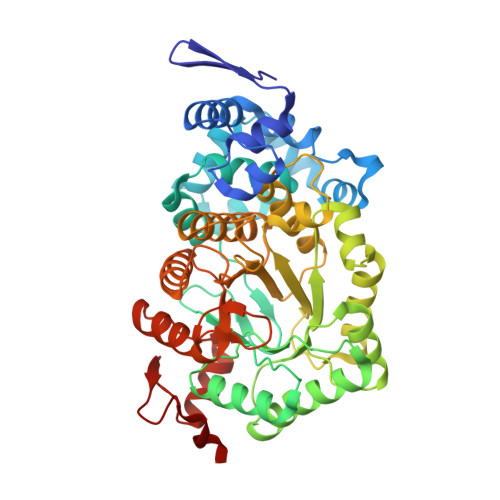
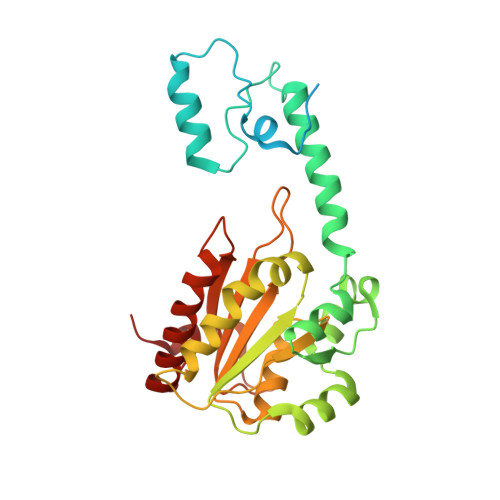
Crystal structures of ethanolamine ammonia-lyase complexed with coenzyme B12 analogs and substrates.
Shibata, N., Tamagaki, H., Hieda, N., Akita, K., Komori, H., Shomura, Y., Terawaki, S., Mori, K., Yasuoka, N., Higuchi, Y., Toraya, T.(2010) J Biological Chem 285: 26484-26493
- PubMed: 20519496
- DOI: https://doi.org/10.1074/jbc.M110.125112
- Primary Citation of Related Structures:
3ABO, 3ABQ, 3ABR, 3ABS - PubMed Abstract:
N-terminal truncation of the Escherichia coli ethanolamine ammonia-lyase beta-subunit does not affect the catalytic properties of the enzyme (Akita, K., Hieda, N., Baba, N., Kawaguchi, S., Sakamoto, H., Nakanishi, Y., Yamanishi, M., Mori, K., and Toraya, T. (2010) J. Biochem. 147, 83-93). The binary complex of the truncated enzyme with cyanocobalamin and the ternary complex with cyanocobalamin or adeninylpentylcobalamin and substrates were crystallized, and their x-ray structures were analyzed. The enzyme exists as a trimer of the (alphabeta)(2) dimer. The active site is in the (beta/alpha)(8) barrel of the alpha-subunit; the beta-subunit covers the lower part of the cobalamin that is bound in the interface of the alpha- and beta-subunits. The structure complexed with adeninylpentylcobalamin revealed the presence of an adenine ring-binding pocket in the enzyme that accommodates the adenine moiety through a hydrogen bond network. The substrate is bound by six hydrogen bonds with active-site residues. Argalpha(160) contributes to substrate binding most likely by hydrogen bonding with the O1 atom. The modeling study implies that marked angular strains and tensile forces induced by tight enzyme-coenzyme interactions are responsible for breaking the coenzyme Co-C bond. The coenzyme adenosyl radical in the productive conformation was modeled by superimposing its adenine ring on the adenine ring-binding site followed by ribosyl rotation around the N-glycosidic bond. A major structural change upon substrate binding was not observed with this particular enzyme. Glualpha(287), one of the substrate-binding residues, has a direct contact with the ribose group of the modeled adenosylcobalamin, which may contribute to the substrate-induced additional labilization of the Co-C bond.
Organizational Affiliation:
Department of Life Science, Graduate School of Life Science, University of Hyogo, Hyogo, Japan. shibach@sci.u-hyogo.ac.jp