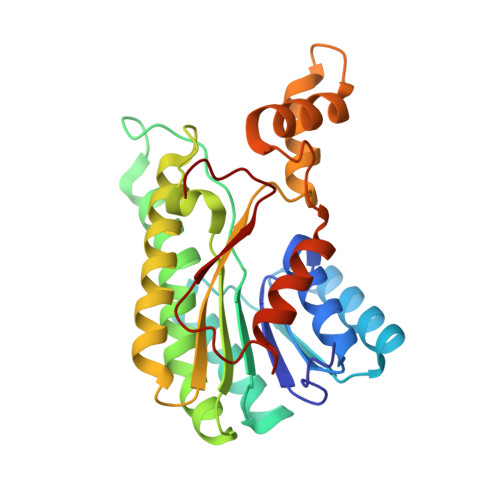
The Crystal Structure of l-Sorbose Reductase from Gluconobacter frateurii Complexed with NADPH and l-Sorbose
Kubota, K., Nagata, K., Okai, M., Miyazono, K., Soemphol, W., Ohtsuka, J., Yamamura, A., Saichana, N., Toyama, H., Matsushita, K., Tanokura, M.(2011) J Mol Biol 407: 543-555
- PubMed: 21277857
- DOI: https://doi.org/10.1016/j.jmb.2011.01.008
- Primary Citation of Related Structures:
3AI1, 3AI2, 3AI3 - PubMed Abstract:
l-Sorbose reductase from Gluconobacter frateurii (SR) is an NADPH-dependent oxidoreductase. SR preferentially catalyzes the reversible reaction between d-sorbitol and l-sorbose with high substrate specificity. To elucidate the structural basis of the catalytic mechanism and the substrate specificity of SR, we have determined the structures of apo-SR, SR in complex with NADPH, and the inactive mutant (His116Leu) of SR in complex with NADPH and l-sorbose at 2.83 Å, 1.90 Å, and 1.80 Å resolutions, respectively. Our results show that SR belongs to the short-chain dehydrogenase/reductase (SDR) family and forms a tetrameric structure. Although His116 is not conserved among SDR family enzymes, the structures of SR have revealed that His116 is important for the stabilization of the proton relay system and for active-site conformation as a fourth catalytic residue. In the ternary complex structure, l-sorbose is recognized by 11 hydrogen bonds. Site-directed mutagenesis of residues around the l-sorbose-binding site has shown that the loss of almost full enzymatic activity was caused by not only the substitution of putative catalytic residues but also the substitution of the residue used for the recognition of the C4 hydroxyl groups of l-sorbose (Glu154) and of the residues used for the construction of the substrate-binding pocket (Cys146 and Gly188). The recognition of the C4 hydroxyl group of l-sorbose would be indispensable for the substrate specificity of SR, which recognizes only l-sorbose and d-sorbitol but not other sugars. Our results indicated that these residues were crucial for the substrate recognition and specificity of SR.
Organizational Affiliation:
Department of Applied Biological Chemistry, Graduate School of Agricultural and Life Sciences, University of Tokyo, 1-1-1 Yayoi, Bunkyo-ku, Tokyo 113-8657, Japan.