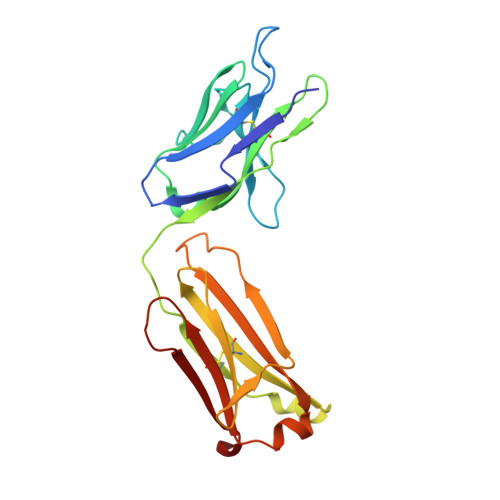
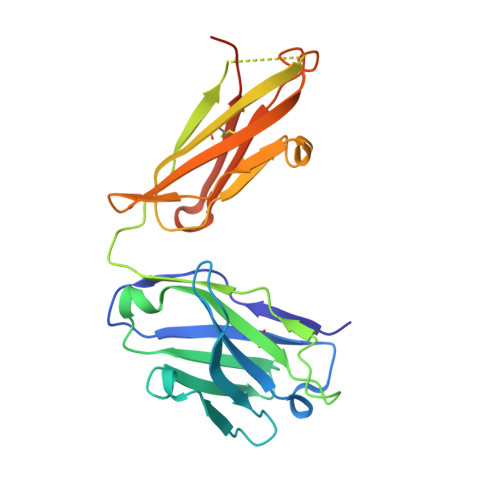
A pipeline for the production of antibody fragments for structural studies using transient expression in HEK 293T cells.
Nettleship, J.E., Ren, J., Rahman, N., Berrow, N.S., Hatherley, D., Barclay, A.N., Owens, R.J.(2008) Protein Expr Purif 62: 83-89
- PubMed: 18662785
- DOI: https://doi.org/10.1016/j.pep.2008.06.017
- Primary Citation of Related Structures:
3DGG, 3DIF - PubMed Abstract:
We describe a pipeline for the rapid production of recombinant Fabs derived from mouse monoclonal antibodies suitable for use in structural studies. The pipeline is exemplified by the production of three Fabs derived from the monoclonal antibodies OX108 (anti-CD200 receptor), OX117 and OX119 (anti-SIRPgamma). Heavy and light chain variable domains were inserted into separate expression vectors containing resident constant regions using In-Fusion PCR cloning. Following transient co-expression in HEK 293T cells, secreted Fab fragments were purified by metal chelate chromatography and gel filtration using an automated procedure with yields of up to 4mg/L of cell culture. Following crystallization trials, diffracting crystals were obtained for the recombinant Fabs of OX108 and OX117, and their structures solved to 2.3A and 2.4A, respectively.
Organizational Affiliation:
The Oxford Protein Production Facility, Henry Wellcome Building for Genomic Medicine, University of Oxford, Oxford, UK. ray@strubi.ox.ac.uk