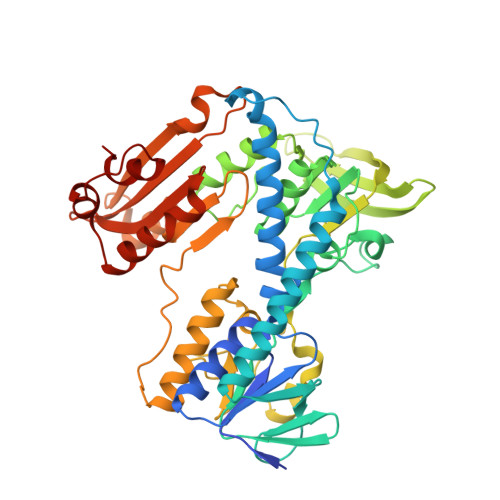
Catalytic cycle of human glutathione reductase near 1 A resolution.
Berkholz, D.S., Faber, H.R., Savvides, S.N., Karplus, P.A.(2008) J Mol Biol 382: 371-384
- PubMed: 18638483
- DOI: https://doi.org/10.1016/j.jmb.2008.06.083
- Primary Citation of Related Structures:
3DJG, 3DJJ, 3DK4, 3DK8, 3DK9 - PubMed Abstract:
Efficient enzyme catalysis depends on exquisite details of structure beyond those resolvable in typical medium- and high-resolution crystallographic analyses. Here we report synchrotron-based cryocrystallographic studies of natural substrate complexes of the flavoenzyme human glutathione reductase (GR) at nominal resolutions between 1.1 and 0.95 A that reveal new aspects of its mechanism. Compression in the active site causes overlapping van der Waals radii and distortion in the nicotinamide ring of the NADPH substrate, which enhances catalysis via stereoelectronic effects. The bound NADPH and redox-active disulfide are positioned optimally on opposite sides of the flavin for a 1,2-addition across a flavin double bond. The new structures extend earlier observations to reveal that the redox-active disulfide loop in GR is an extreme case of sequential peptide bonds systematically deviating from planarity--a net deviation of 53 degrees across five residues. But this apparent strain is not a factor in catalysis, as it is present in both oxidized and reduced structures. Intriguingly, the flavin bond lengths in oxidized GR are intermediate between those expected for oxidized and reduced flavin, but we present evidence that this may not be due to the protein environment but instead due to partial synchrotron reduction of the flavin by the synchrotron beam. Finally, of more general relevance, we present evidence that the structures of synchrotron-reduced disulfide bonds cannot generally be used as reliable models for naturally reduced disulfide bonds.
Organizational Affiliation:
Department of Biochemistry and Biophysics, Oregon State University, 2011 ALS, Corvallis, OR 97331-7305, USA.