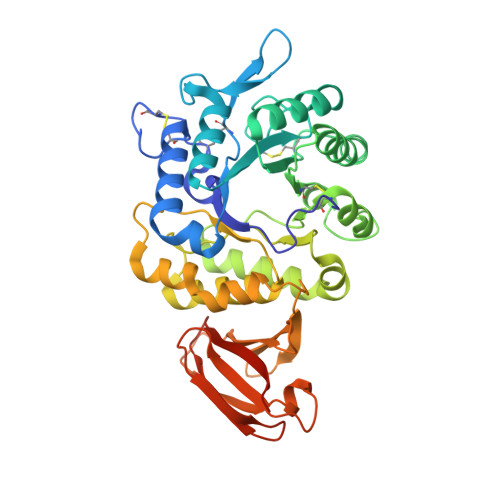
The 1.9 a structure of human alpha-N-acetylgalactosaminidase: The molecular basis of Schindler and Kanzaki diseases
Clark, N.E., Garman, S.C.(2009) J Mol Biol 393: 435-447
- PubMed: 19683538
- DOI: https://doi.org/10.1016/j.jmb.2009.08.021
- Primary Citation of Related Structures:
3H53, 3H54, 3H55, 3IGU - PubMed Abstract:
alpha-N-acetylgalactosaminidase (alpha-NAGAL; E.C. 3.2.1.49) is a lysosomal exoglycosidase that cleaves terminal alpha-N-acetylgalactosamine residues from glycopeptides and glycolipids. In humans, a deficiency of alpha-NAGAL activity results in the lysosomal storage disorders Schindler disease and Kanzaki disease. To better understand the molecular defects in the diseases, we determined the crystal structure of human alpha-NAGAL after expressing wild-type and glycosylation-deficient glycoproteins in recombinant insect cell expression systems. We measured the enzymatic parameters of our purified wild-type and mutant enzymes, establishing their enzymatic equivalence. To investigate the binding specificity and catalytic mechanism of the human alpha-NAGAL enzyme, we determined three crystallographic complexes with different catalytic products bound in the active site of the enzyme. To better understand how individual defects in the alpha-NAGAL glycoprotein lead to Schindler disease, we analyzed the effect of disease-causing mutations on the three-dimensional structure.
Organizational Affiliation:
Department of Biochemistry and Molecular Biology, University of Massachusetts, Amherst, 01003, USA.