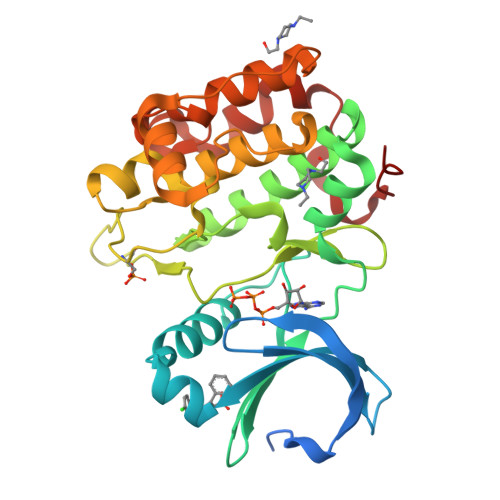
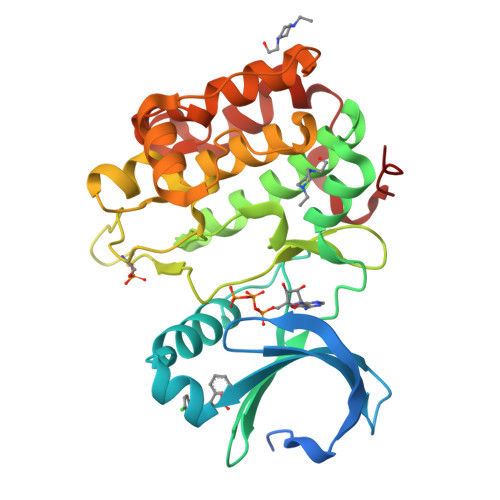
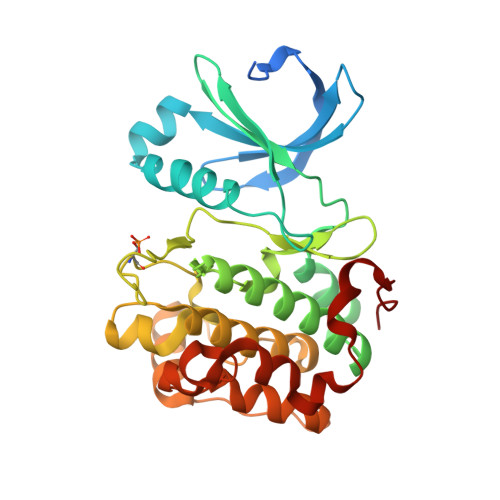
Structure and allosteric effects of low-molecular-weight activators on the protein kinase PDK1.
Hindie, V., Stroba, A., Zhang, H., Lopez-Garcia, L.A., Idrissova, L., Zeuzem, S., Hirschberg, D., Schaeffer, F., Jrgensen, T.J., Engel, M., Alzari, P.M., Biondi, R.M.(2009) Nat Chem Biol 5: 758-764
- PubMed: 19718043
- DOI: https://doi.org/10.1038/nchembio.208
- Primary Citation of Related Structures:
3HRC, 3HRF - PubMed Abstract:
Protein phosphorylation transduces a large set of intracellular signals. One mechanism by which phosphorylation mediates signal transduction is by prompting conformational changes in the target protein or interacting proteins. Previous work described an allosteric site mediating phosphorylation-dependent activation of AGC kinases. The AGC kinase PDK1 is activated by the docking of a phosphorylated motif from substrates. Here we present the crystallography of PDK1 bound to a rationally developed low-molecular-weight activator and describe the conformational changes induced by small compounds in the crystal and in solution using a fluorescence-based assay and deuterium exchange experiments. Our results indicate that the binding of the compound produces local changes at the target site, the PIF binding pocket, and also allosteric changes at the ATP binding site and the activation loop. Altogether, we present molecular details of the allosteric changes induced by small compounds that trigger the activation of PDK1 through mimicry of phosphorylation-dependent conformational changes.
Organizational Affiliation:
Research Group PhosphoSites, Department of Internal Medicine I, Universitätsklinikum Frankfurt, Frankfurt, Germany.