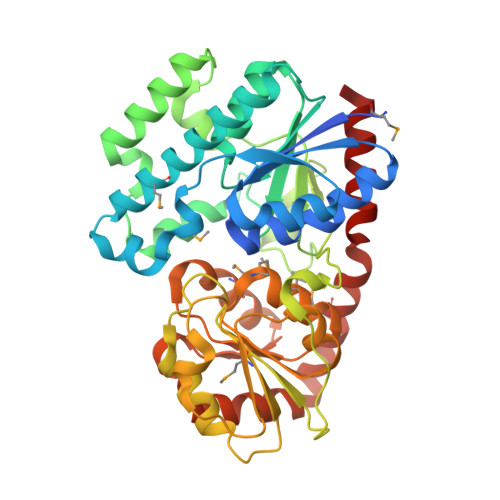
Complete set of glycosyltransferase structures in the calicheamicin biosynthetic pathway reveals the origin of regiospecificity.
Chang, A., Singh, S., Helmich, K.E., Goff, R.D., Bingman, C.A., Thorson, J.S., Phillips, G.N.(2011) Proc Natl Acad Sci U S A 108: 17649-17654
- PubMed: 21987796
- DOI: https://doi.org/10.1073/pnas.1108484108
- Primary Citation of Related Structures:
3IA7, 3IAA, 3OTG, 3OTH, 3OTI, 3RSC - PubMed Abstract:
Glycosyltransferases are useful synthetic catalysts for generating natural products with sugar moieties. Although several natural product glycosyltransferase structures have been reported, design principles of glycosyltransferase engineering for the generation of glycodiversified natural products has fallen short of its promise, partly due to a lack of understanding of the relationship between structure and function. Here, we report structures of all four calicheamicin glycosyltransferases (CalG1, CalG2, CalG3, and CalG4), whose catalytic functions are clearly regiospecific. Comparison of these four structures reveals a conserved sugar donor binding motif and the principles of acceptor binding region reshaping. Among them, CalG2 possesses a unique catalytic motif for glycosylation of hydroxylamine. Multiple glycosyltransferase structures in a single natural product biosynthetic pathway are a valuable resource for understanding regiospecific reactions and substrate selectivities and will help future glycosyltransferase engineering.
Organizational Affiliation:
Department of Biochemistry, University of Wisconsin, 433 Babcock Drive, Madison, WI 53706, USA.