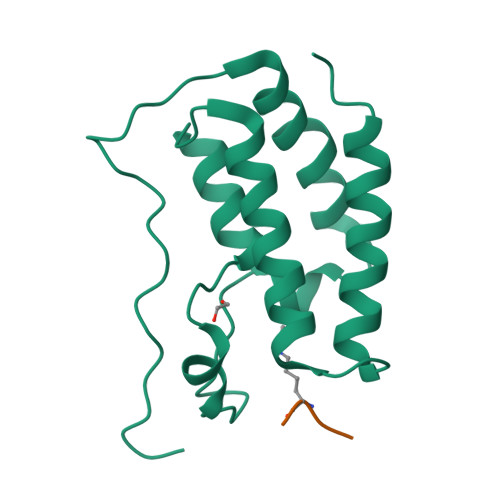
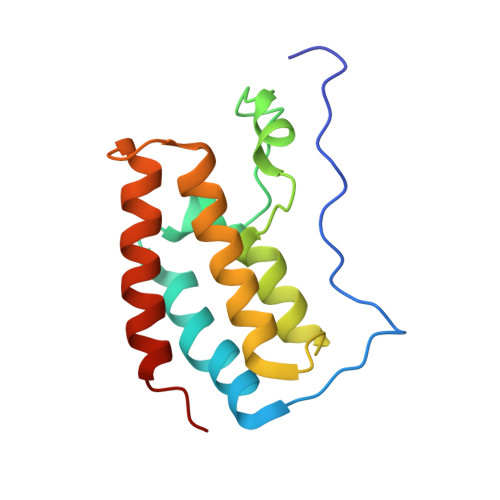
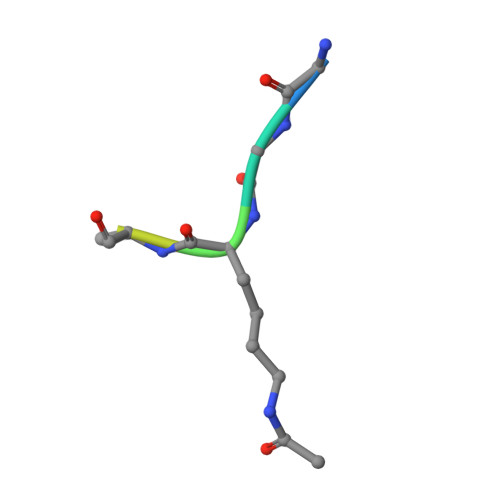
Structures of the Dual Bromodomains of the P-TEFb-activating Protein Brd4 at Atomic Resolution
Vollmuth, F., Blankenfeldt, W., Geyer, M.(2009) J Biological Chem 284: 36547-36556
- PubMed: 19828451
- DOI: https://doi.org/10.1074/jbc.M109.033712
- Primary Citation of Related Structures:
3JVJ, 3JVK, 3JVL, 3JVM - PubMed Abstract:
Brd4 is a member of the bromodomains and extra terminal domain (BET) family of proteins that recognize acetylated chromatin structures through their bromodomains and act as transcriptional activators. Brd4 functions as an associated factor and positive regulator of P-TEFb, a Cdk9-cyclin T heterodimer that stimulates transcriptional elongation by RNA polymerase II. Here, the crystal structures of the two bromodomains of Brd4 (BD1 and BD2) were determined at 1.5 and 1.2 A resolution, respectively. Complex formation of BD1 with a histone H3 tail polypeptide encompassing residues 12-19 showed binding of the Nzeta-acetylated lysine 14 to the conserved asparagine 140 of Brd4. In contrast, in BD2 the N-terminal linker sequence was found to interact with the binding site for acetylated lysines of the adjacent molecule to form continuous strings in the crystal lattice. This assembly shows for the first time a different binding ligand than acetylated lysine indicating that also other sequence compositions may be able to form similar interaction networks. Isothermal titration calorimetry revealed best binding of BD1 to H3 and of BD2 to H4 acetylated lysine sequences, suggesting alternating histone recognition specificities. Intriguingly, an acetylated lysine motif from cyclin T1 bound similarly well to BD2. Whereas the structure of Brd2 BD1 suggested its dimer formation, both Brd4 bromodomains appeared monomeric in solution as shown by size exclusion chromatography and mutational analyses.
Organizational Affiliation:
Abteilung Physikalische Biochemie, Max-Planck-Institut für Molekulare Physiologie, Otto-Hahn-Strasse 11, D-44227 Dortmund, Germany.