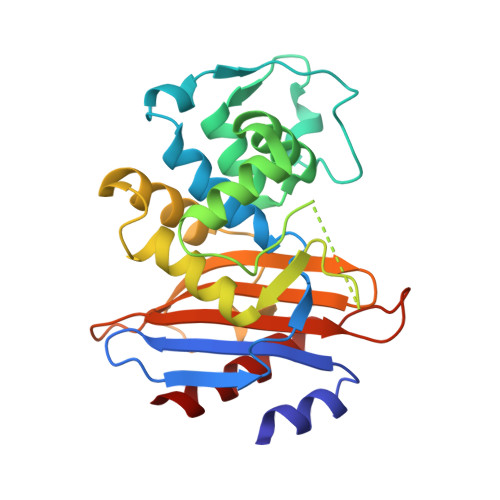
Structural studies of the mechanism for biosensing antibiotics in a fluorescein-labeled beta-lactamase.
Wong, W.T., Au, H.W., Yap, H.K., Leung, Y.C., Wong, K.Y., Zhao, Y.(2011) BMC Struct Biol 11: 15-15
- PubMed: 21443768
- DOI: https://doi.org/10.1186/1472-6807-11-15
- Primary Citation of Related Structures:
3M2K - PubMed Abstract:
β-lactamase conjugated with environment-sensitive fluorescein molecule to residue 166 on the Ω-loop near its catalytic site is a highly effective biosensor for β-lactam antibiotics. Yet the molecular mechanism of such fluorescence-based biosensing is not well understood. Here we report the crystal structure of a Class A β-lactamase PenP from Bacillus licheniformis 749/C with fluorescein conjugated at residue 166 after E166C mutation, both in apo form (PenP-E166Cf) and in covalent complex form with cefotaxime (PenP-E166Cf-cefotaxime), to illustrate its biosensing mechanism. In the apo structure the fluorescein molecule partially occupies the antibiotic binding site and is highly dynamic. In the PenP-E166Cf-cefatoxime complex structure the binding and subsequent acylation of cefotaxime to PenP displaces fluorescein from its original location to avoid steric clash. Such displacement causes the well-folded Ω-loop to become fully flexible and the conjugated fluorescein molecule to relocate to a more solvent exposed environment, hence enhancing its fluorescence emission. Furthermore, the fully flexible Ω-loop enables the narrow-spectrum PenP enzyme to bind cefotaxime in a mode that resembles the extended-spectrum β-lactamase. Our structural studies indicate the biosensing mechanism of a fluorescein-labelled β-lactamase. Such findings confirm our previous proposal based on molecular modelling and provide useful information for the rational design of β-lactamase-based biosensor to detect the wide spectrum of β-lactam antibiotics. The observation of increased Ω-loop flexibility upon conjugation of fluorophore may have the potential to serve as a screening tool for novel β-lactamase inhibitors that target the Ω-loop and not the active site.
Organizational Affiliation:
Department of Applied Biology and Chemical Technology, Central Laboratory of Institute of Molecular Technology for Drug Discovery and Synthesis, The Hong Kong Polytechnic University, Hung Hom, Hong Hong, China.