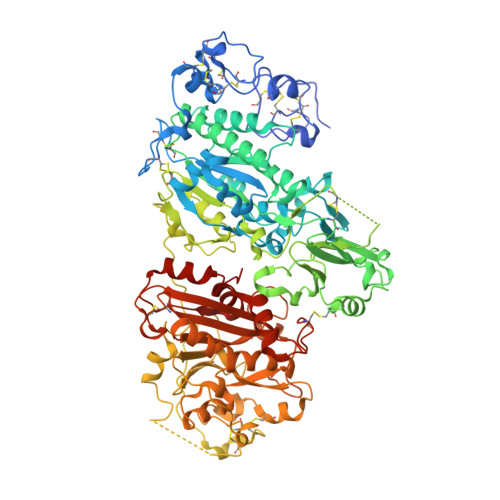
Crystal structure of autotaxin and insight into GPCR activation by lipid mediators
Nishimasu, H., Okudaira, S., Hama, K., Mihara, E., Dohmae, N., Inoue, A., Ishitani, R., Takagi, J., Aoki, J., Nureki, O.(2011) Nat Struct Mol Biol 18: 205-212
- PubMed: 21240269
- DOI: https://doi.org/10.1038/nsmb.1998
- Primary Citation of Related Structures:
3NKM, 3NKN, 3NKO, 3NKP, 3NKQ, 3NKR - PubMed Abstract:
Autotaxin (ATX, also known as Enpp2) is a secreted lysophospholipase D that hydrolyzes lysophosphatidylcholine to generate lysophosphatidic acid (LPA), a lipid mediator that activates G protein-coupled receptors to evoke various cellular responses. Here, we report the crystal structures of mouse ATX alone and in complex with LPAs with different acyl-chain lengths and saturations. These structures reveal that the multidomain architecture helps to maintain the structural rigidity of the lipid-binding pocket, which accommodates the respective LPA molecules in distinct conformations. They indicate that a loop region in the catalytic domain is a major determinant for the substrate specificity of the Enpp family enzymes. Furthermore, along with biochemical and biological data, these structures suggest that the produced LPAs are delivered from the active site to cognate G protein-coupled receptors through a hydrophobic channel.
Organizational Affiliation:
Department of Biophysics and Biochemistry, Graduate School of Science, The University of Tokyo, Tokyo, Japan.