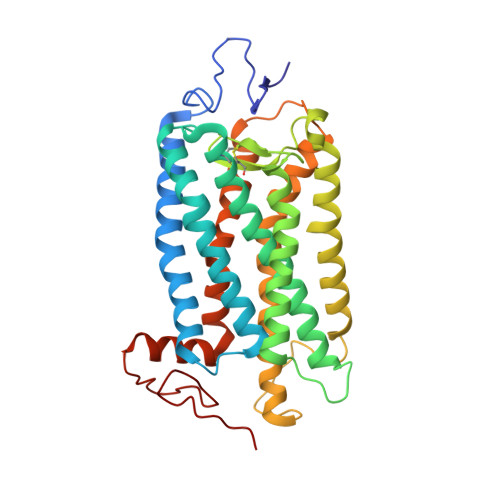
Binding of more than one retinoid to visual opsins
Makino, C.L., Riley, C.K., Looney, J., Crouch, R.K., Okada, T.(2010) Biophys J 99: 2366-2373
- PubMed: 20923672
- DOI: https://doi.org/10.1016/j.bpj.2010.08.003
- Primary Citation of Related Structures:
3OAX - PubMed Abstract:
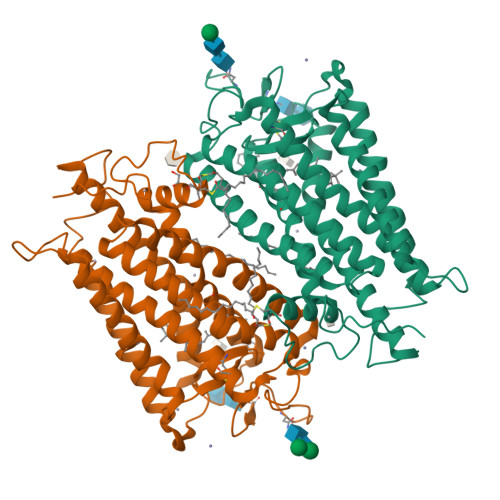
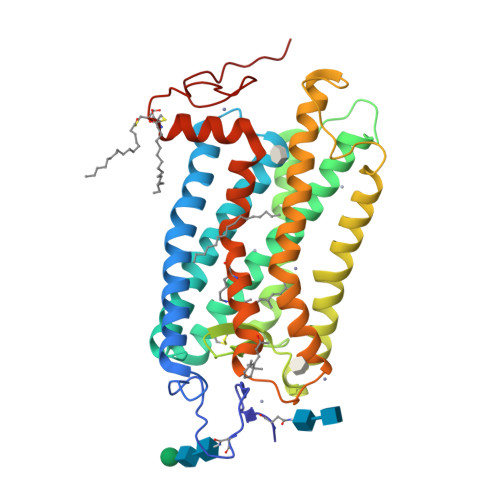
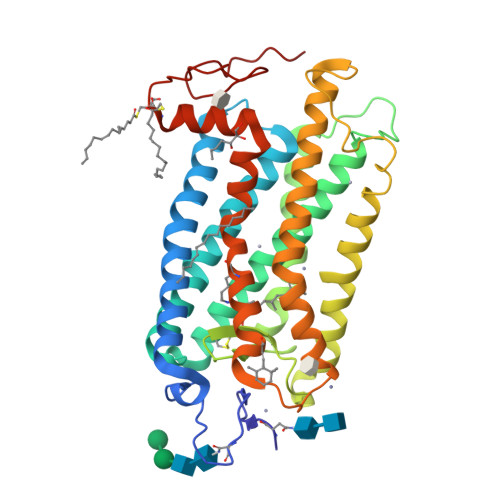
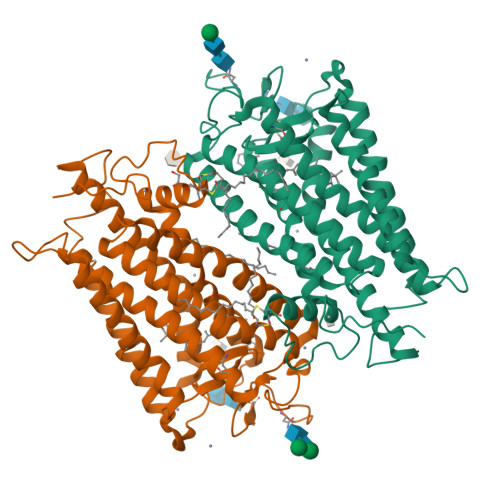
Visual opsins bind 11-cis retinal at an orthosteric site to form rhodopsins but increasing evidence suggests that at least some are capable of binding an additional retinoid(s) at a separate, allosteric site(s). Microspectrophotometric measurements on isolated, dark-adapted, salamander photoreceptors indicated that the truncated retinal analog, β-ionone, partitioned into the membranes of green-sensitive rods; however, in blue-sensitive rod outer segments, there was an enhanced uptake of four or more β-ionones per rhodopsin. X-ray crystallography revealed binding of one β-ionone to bovine green-sensitive rod rhodopsin. Cocrystallization only succeeded with extremely high concentrations of β-ionone and binding did not alter the structure of rhodopsin from the inactive state. Salamander green-sensitive rod rhodopsin is also expected to bind β-ionone at sufficiently high concentrations because the binding site is present on its surface. Therefore, both blue- and green-sensitive rod rhodopsins have at least one allosteric binding site for retinoid, but β-ionone binds to the latter type of rhodopsin with low affinity and low efficacy.
Organizational Affiliation:
Department of Ophthalmology, Massachusetts Eye and Ear Infirmary, Boston, MA, USA. cmakino@meei.harvard.edu