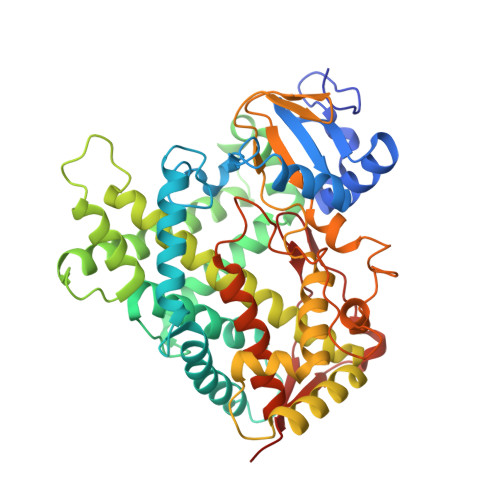
Structural comparison of cytochromes P450 2A6, 2A13, and 2E1 with pilocarpine.
DeVore, N.M., Meneely, K.M., Bart, A.G., Stephens, E.S., Battaile, K.P., Scott, E.E.(2012) FEBS J 279: 1621-1631
- PubMed: 22051186
- DOI: https://doi.org/10.1111/j.1742-4658.2011.08412.x
- Primary Citation of Related Structures:
3T3Q, 3T3R, 3T3S, 3T3Z - PubMed Abstract:
Human xenobiotic-metabolizing cytochrome P450 (CYP) enzymes can each bind and monooxygenate a diverse set of substrates, including drugs, often producing a variety of metabolites. Additionally, a single ligand can interact with multiple CYP enzymes, but often the protein structural similarities and differences that mediate such overlapping selectivity are not well understood. Even though the CYP superfamily has a highly canonical global protein fold, there are large variations in the active site size, topology, and conformational flexibility. We have determined how a related set of three human CYP enzymes bind and interact with a common inhibitor, the muscarinic receptor agonist drug pilocarpine. Pilocarpine binds and inhibits the hepatic CYP2A6 and respiratory CYP2A13 enzymes much more efficiently than the hepatic CYP2E1 enzyme. To elucidate key residues involved in pilocarpine binding, crystal structures of CYP2A6 (2.4 Å), CYP2A13 (3.0 Å), CYP2E1 (2.35 Å), and the CYP2A6 mutant enzyme, CYP2A6 I208S/I300F/G301A/S369G (2.1 Å) have been determined with pilocarpine in the active site. In all four structures, pilocarpine coordinates to the heme iron, but comparisons reveal how individual residues lining the active sites of these three distinct human enzymes interact differently with the inhibitor pilocarpine.
Organizational Affiliation:
Department of Medicinal Chemistry, The University of Kansas, Lawrence, KS 66045, USA.