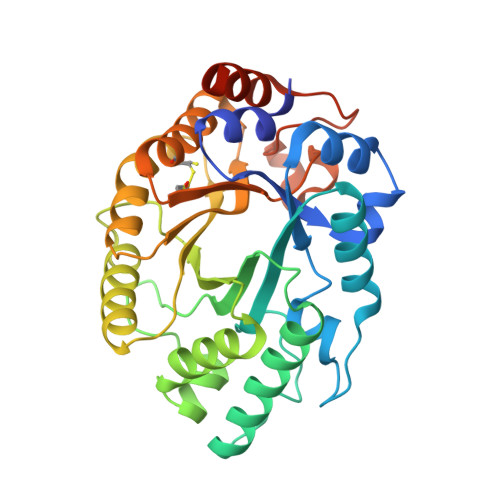
Precision is Essential for Efficient Catalysis in an Evolved Kemp Eliminase
Blomberg, R., Kries, H., Pinkas, D.M., Mittl, P.R.E., Gruetter, M.G., Privett, H.K., Mayo, S.L., Hilvert, D.(2013) Nature 503: 418
- PubMed: 24132235
- DOI: https://doi.org/10.1038/nature12623
- Primary Citation of Related Structures:
4BS0 - PubMed Abstract:
Linus Pauling established the conceptual framework for understanding and mimicking enzymes more than six decades ago. The notion that enzymes selectively stabilize the rate-limiting transition state of the catalysed reaction relative to the bound ground state reduces the problem of design to one of molecular recognition. Nevertheless, past attempts to capitalize on this idea, for example by using transition state analogues to elicit antibodies with catalytic activities, have generally failed to deliver true enzymatic rates. The advent of computational design approaches, combined with directed evolution, has provided an opportunity to revisit this problem. Starting from a computationally designed catalyst for the Kemp elimination--a well-studied model system for proton transfer from carbon--we show that an artificial enzyme can be evolved that accelerates an elementary chemical reaction 6 × 10(8)-fold, approaching the exceptional efficiency of highly optimized natural enzymes such as triosephosphate isomerase. A 1.09 Å resolution crystal structure of the evolved enzyme indicates that familiar catalytic strategies such as shape complementarity and precisely placed catalytic groups can be successfully harnessed to afford such high rate accelerations, making us optimistic about the prospects of designing more sophisticated catalysts.
Organizational Affiliation:
1] Laboratory of Organic Chemistry, ETH Zurich, 8093 Zurich, Switzerland [2] Corporate RD Division, Firmenich SA, 1211 Geneva, Switzerland (R.B.); Protabit, Pasadena, California 91101, USA (H.K.P.).