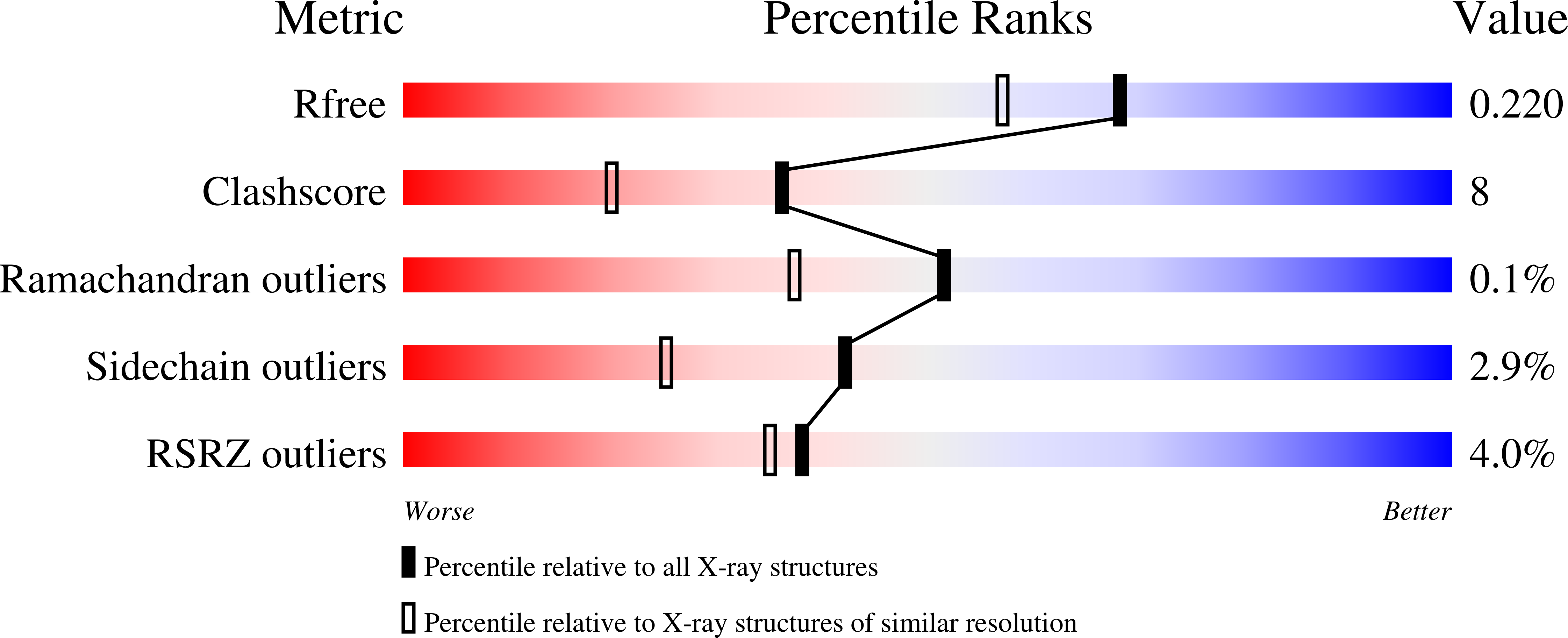
VFV as a New Effective CYP51 Structure-Derived Drug Candidate for Chagas Disease and Visceral Leishmaniasis.
Lepesheva, G.I., Hargrove, T.Y., Rachakonda, G., Wawrzak, Z., Pomel, S., Cojean, S., Nde, P.N., Nes, W.D., Locuson, C.W., Calcutt, M.W., Waterman, M.R., Daniels, J.S., Loiseau, P.M., Villalta, F.(2015) J Infect Dis 212: 1439-1448
- PubMed: 25883390
- DOI: https://doi.org/10.1093/infdis/jiv228
- Primary Citation of Related Structures:
4G3J, 4G7G - PubMed Abstract:
Sterol 14α-demethylases (CYP51) are the enzymes essential for sterol biosynthesis. They serve as clinical targets for antifungal azoles and are considered as targets for treatment of human Trypanosomatidae infections. Recently, we have shown that VNI, a potent and selective inhibitor of trypanosomal CYP51 that we identified and structurally characterized in complex with the enzyme, can cure the acute and chronic forms of Chagas disease. The purpose of this work was to apply the CYP51 structure/function for further development of the VNI scaffold. As anticipated, VFV (R)-N-(1-(3,4'-difluorobiphenyl-4-yl)-2-(1H-imidazol-1-yl)ethyl)-4-(5-phenyl-1,3,4-oxadiazol-2-yl)benzamide, the derivative designed to fill the deepest portion of the CYP51 substrate-binding cavity, reveals a broader antiprotozoan spectrum of action. It has stronger antiparasitic activity in cellular experiments, cures the experimental Chagas disease with 100% efficacy, and suppresses visceral leishmaniasis by 89% (vs 60% for VNI). Oral bioavailability, low off-target activity, favorable pharmacokinetics and tissue distribution characterize VFV as a promising new drug candidate.
Organizational Affiliation:
Department of Biochemistry School of Medicine Center for Structural Biology, Vanderbilt University.