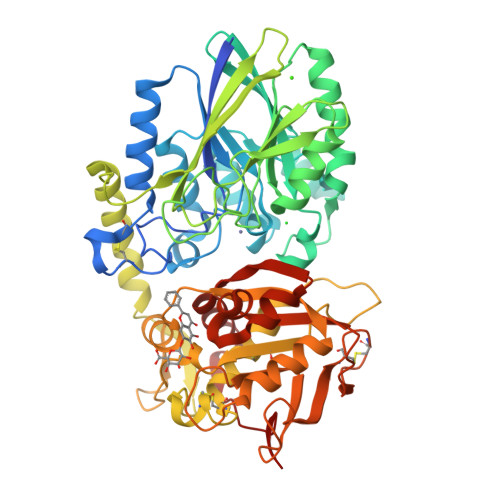
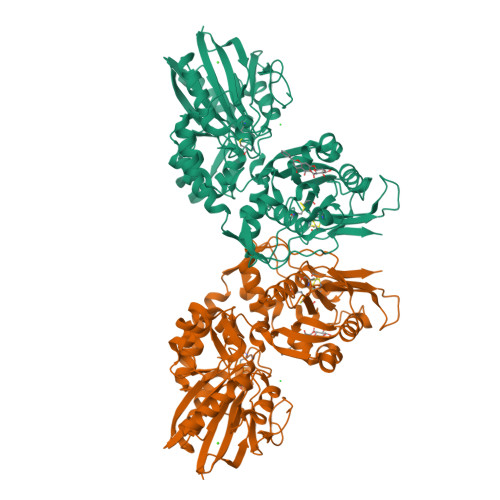
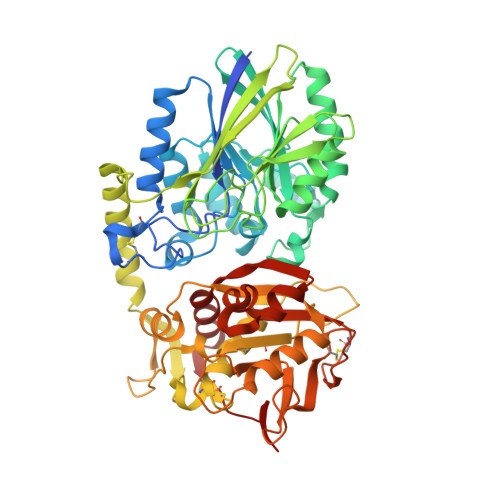
Crystal Structure of the Human Ecto-5'-Nucleotidase (CD73): Insights into the Regulation of Purinergic Signaling.
Knapp, K., Zebisch, M., Pippel, J., El-Tayeb, A., Muller, C.E., Strater, N.(2012) Structure 20: 2161-2173
- PubMed: 23142347
- DOI: https://doi.org/10.1016/j.str.2012.10.001
- Primary Citation of Related Structures:
4H1Y, 4H2B, 4H2F, 4H2G, 4H2I - PubMed Abstract:
In vertebrates ecto-5'-nucleotidase (e5NT) catalyzes the hydrolysis of extracellular AMP to adenosine and represents the major control point for extracellular adenosine levels. Due to its pivotal role for activation of P1 adenosine receptors, e5NT has emerged as an appealing drug target for treatment of inflammation, chronic pain, hypoxia, and cancer. Crystal structures of the dimeric human e5NT reveal an extensive 114° conformational switch between the open and closed forms of the enzyme. The dimerization interface is formed by the C-terminal domains and exhibits interchain motions of up to 13°. Complex structures with adenosine and AMPCP indicate that structural control of the domain movement determines the selectivity for monophosphate nucleotides. Binding modes of nucleotide-derived and flavonoid-based compounds complexed to the C-terminal domain in the open form reveal an additional binding pocket of ∼210 Å(3) that might be explored to design more potent inhibitors.
Organizational Affiliation:
Institute of Bioanalytical Chemistry, Center for Biotechnology and Biomedicine, University of Leipzig, Deutscher Platz 5, 04103 Leipzig, Germany.