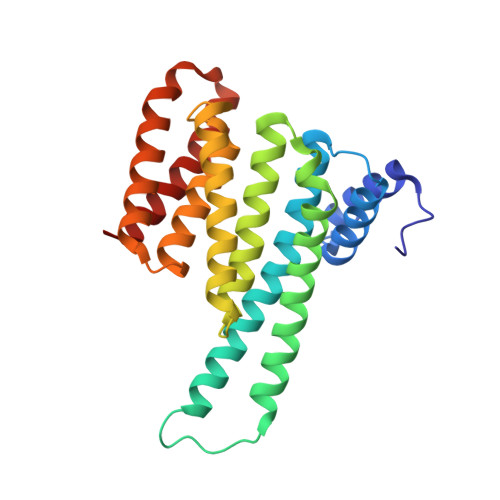
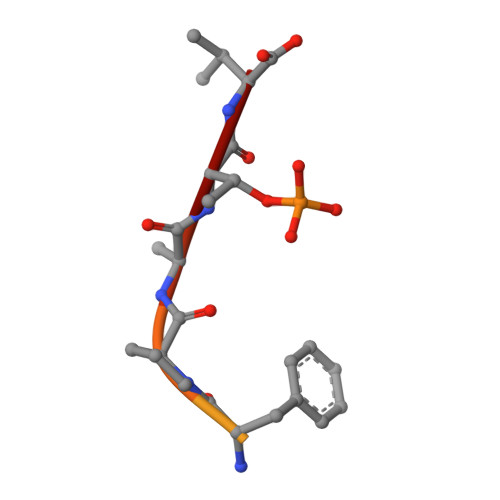
Interaction of 14-3-3 proteins with the estrogen receptor alpha F domain provides a drug target interface.
De Vries-van Leeuwen, I.J., da Costa Pereira, D., Flach, K.D., Piersma, S.R., Haase, C., Bier, D., Yalcin, Z., Michalides, R., Feenstra, K.A., Jimenez, C.R., de Greef, T.F., Brunsveld, L., Ottmann, C., Zwart, W., de Boer, A.H.(2013) Proc Natl Acad Sci U S A 110: 8894-8899
- PubMed: 23676274
- DOI: https://doi.org/10.1073/pnas.1220809110
- Primary Citation of Related Structures:
4JC3, 4JDD - PubMed Abstract:
Estrogen receptor alpha (ERα) is involved in numerous physiological and pathological processes, including breast cancer. Breast cancer therapy is therefore currently directed at inhibiting the transcriptional potency of ERα, either by blocking estrogen production through aromatase inhibitors or antiestrogens that compete for hormone binding. Due to resistance, new treatment modalities are needed and as ERα dimerization is essential for its activity, interference with receptor dimerization offers a new opportunity to exploit in drug design. Here we describe a unique mechanism of how ERα dimerization is negatively controlled by interaction with 14-3-3 proteins at the extreme C terminus of the receptor. Moreover, the small-molecule fusicoccin (FC) stabilizes this ERα/14-3-3 interaction. Cocrystallization of the trimeric ERα/14-3-3/FC complex provides the structural basis for this stabilization and shows the importance of phosphorylation of the penultimate Threonine (ERα-T(594)) for high-affinity interaction. We confirm that T(594) is a distinct ERα phosphorylation site in the breast cancer cell line MCF-7 using a phospho-T(594)-specific antibody and by mass spectrometry. In line with its ERα/14-3-3 interaction stabilizing effect, fusicoccin reduces the estradiol-stimulated ERα dimerization, inhibits ERα/chromatin interactions and downstream gene expression, resulting in decreased cell proliferation. Herewith, a unique functional phosphosite and an alternative regulation mechanism of ERα are provided, together with a small molecule that selectively targets this ERα/14-3-3 interface.
Organizational Affiliation:
Department of Structural Biology, Faculty Earth and Life Sciences, Vrije Universiteit, 1081 HV Amsterdam, The Netherlands.