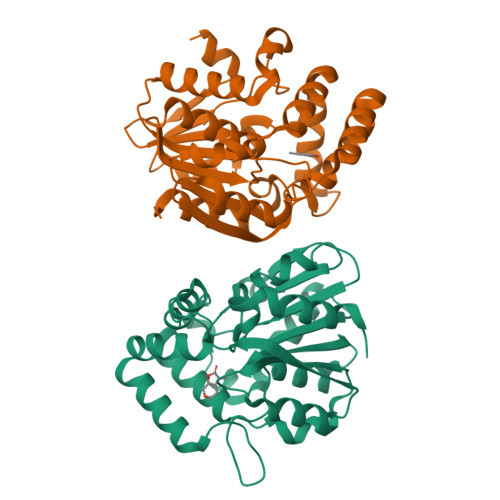
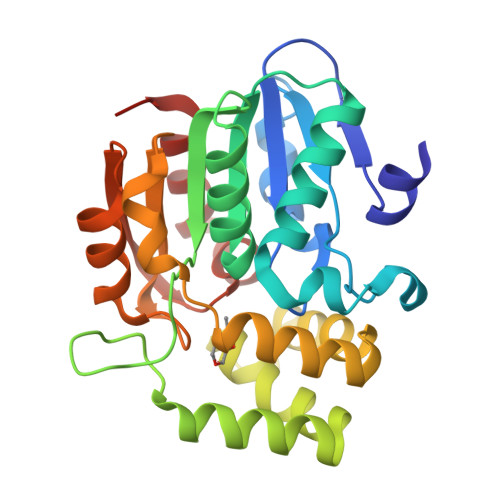
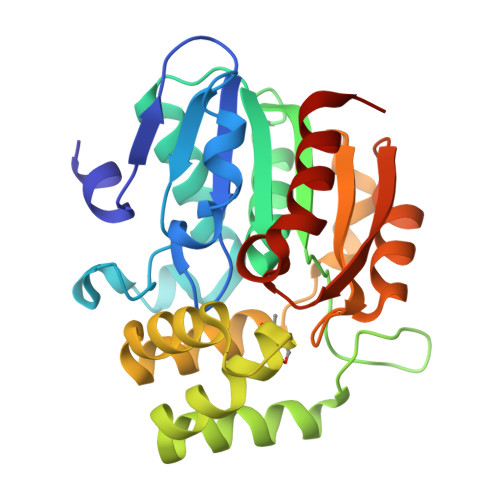
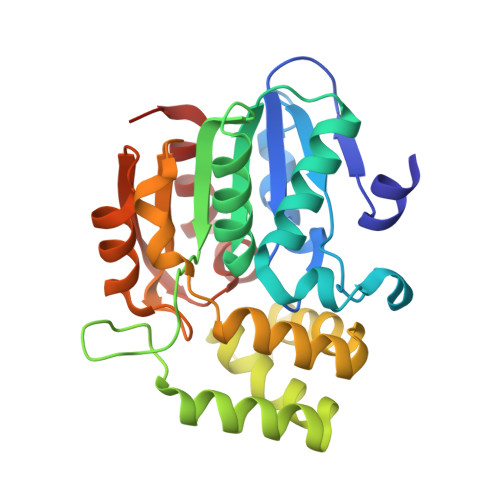
Smoke-derived karrikin perception by the alpha/beta-hydrolase KAI2 from Arabidopsis.
Guo, Y., Zheng, Z., La Clair, J.J., Chory, J., Noel, J.P.(2013) Proc Natl Acad Sci U S A 110: 8284-8289
- PubMed: 23613584
- DOI: https://doi.org/10.1073/pnas.1306265110
- Primary Citation of Related Structures:
4JYM, 4JYP - PubMed Abstract:
Genetic studies in Arabidopsis implicate an α/β-hydrolase, KARRIKIN-INSENSITIVE 2 (KAI2) as a receptor for karrikins, germination-promoting butenolide small molecules found in the smoke of burned plants. However, direct biochemical evidence for the interaction between KAI2 and karrikin and for the mechanism of downstream signaling by a KAI2-karrikin complex remain elusive. We report crystallographic analyses and ligand-binding experiments for KAI2 recognition of karrikins. The karrikin-1 (KAR1) ligand sits in the opening to the active site abutting a helical domain insert but distal from the canonical catalytic triad (Ser95-His246-Asp217) of α/β-hydrolases, consistent with the lack of detectable hydrolytic activity by purified KAI2. The closest approach of KAR1 to Ser95-His246-Asp217 is 3.8 Å from His246. Six aromatic side chains, including His246, encapsulate KAR1 through geometrically defined aromatic-aromatic interactions. KAR1 binding induces a conformational change in KAI2 at the active site entrance. A crevice of hydrophobic residues linking the polar edge of KAR1 and the helical domain insert suggests that KAI2-KAR1 creates a contiguous interface for binding signaling partners in a ligand-dependent manner.
Organizational Affiliation:
Jack H. Skirball Center for Chemical Biology and Proteomics, Salk Institute for Biological Studies, La Jolla, CA 92037, USA.