Azide inhibition of urate oxidase.
Gabison, L., Colloc'h, N., Prange, T.(2014) Acta Crystallogr Sect F Struct Biol Cryst Commun 70: 896-902
- PubMed: 25005084
- DOI: https://doi.org/10.1107/S2053230X14011753
- Primary Citation of Related Structures:
4OQC, 4POE, 4PR8, 4PUV - PubMed Abstract:
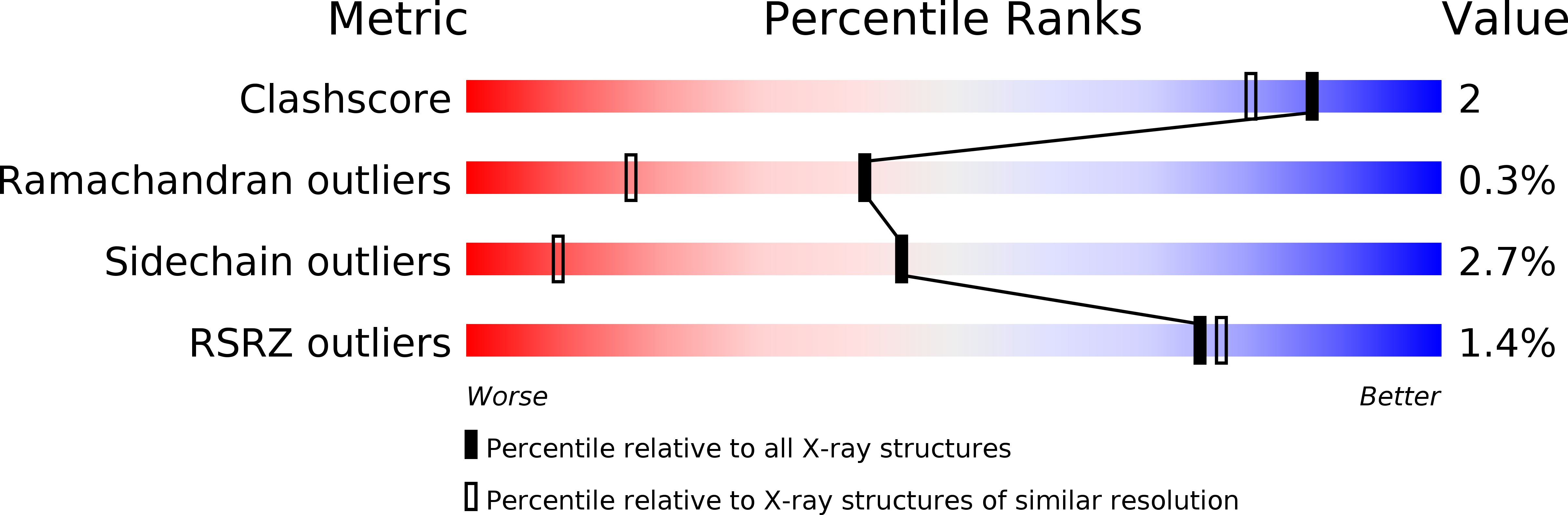
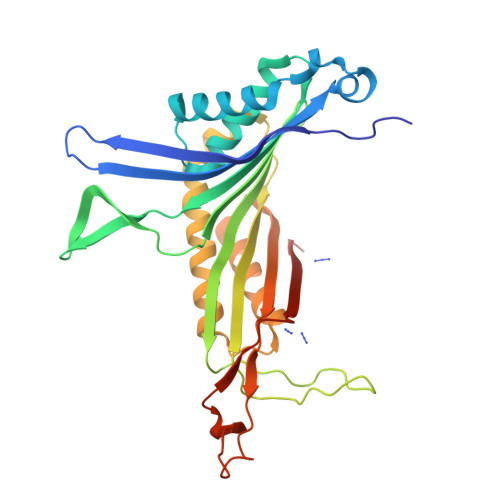
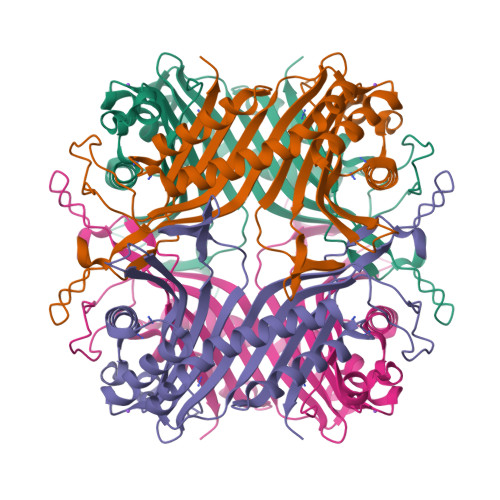
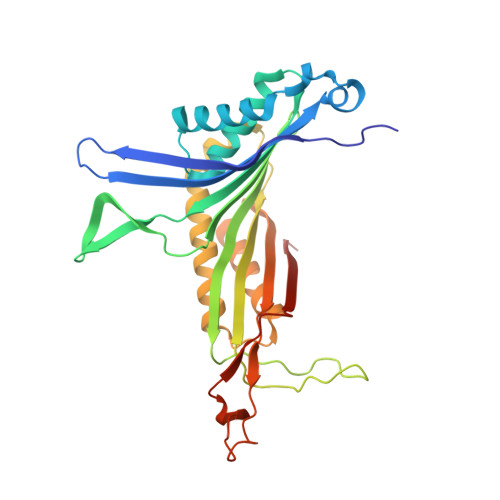
The inhibition of urate oxidase (UOX) by azide was investigated by X-ray diffraction techniques and compared with cyanide inhibition. Two well characterized sites for reagents are present in the enzyme: the dioxygen site and the substrate-binding site. To examine the selectivity of these sites towards azide inhibition, several crystallization conditions were developed. UOX was co-crystallized with azide (N3) in the presence or absence of either uric acid (UA, the natural substrate) or 8-azaxanthine (8AZA, a competitive inhibitor). In a second set of experiments, previously grown orthorhombic crystals of the UOX-UA or UOX-8AZA complexes were soaked in sodium azide solutions. In a third set of experiments, orthorhombic crystals of UOX with the exchangeable ligand 8-nitroxanthine (8NXN) were soaked in a solution containing uric acid and azide simultaneously (competitive soaking). In all assays, the soaking periods were either short (a few hours) or long (one or two months). These different experimental conditions showed that one or other of the sites, or the two sites together, could be inhibited. This also demonstrated that azide not only competes with dioxygen as cyanide does but also competes with the substrate for its enzymatic site. A model in agreement with experimental data would be an azide in equilibrium between two sites, kinetically in favour of the dioxygen site and thermodynamically in favour of the substrate-binding site.
Organizational Affiliation:
Faculty of Pharmacy, UMR 8015 CNRS Laboratoire de Cristallographie et RMN Biologiques, 4 Avenue de l'Observatoire, 75006 Paris, France.