Determinants of Activity at Human Toll-like Receptors 7 and 8: Quantitative Structure-Activity Relationship (QSAR) of Diverse Heterocyclic Scaffolds
Yoo, E., Salunke, D.B., Sil, D., Guo, X., Salyer, A.C.D., Hermanson, A.R., Kumar, M., Malladi, S.S., Balakrishna, R., Thompson, W.H., Tanji, H., Ohto, U., Shimizu, T., David, S.A.(2014) J Med Chem 57: 7955-7970
- PubMed: 25192394
- DOI: https://doi.org/10.1021/jm500744f
- Primary Citation of Related Structures:
4QBZ, 4QC0 - PubMed Abstract:
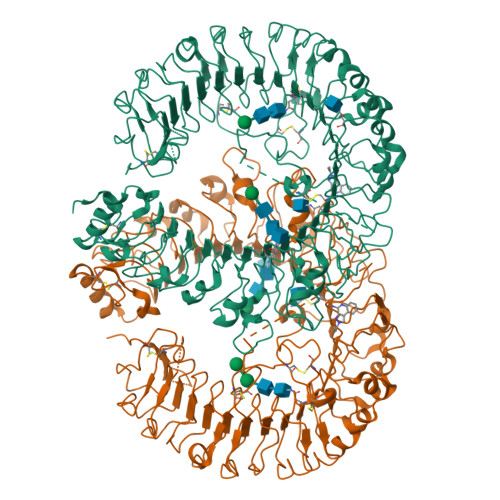
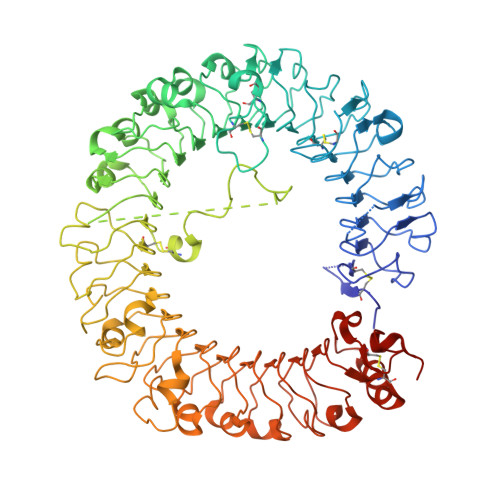
Toll-like receptor (TLR) 7 and 8 agonists are potential vaccine adjuvants, since they directly activate APCs and enhance Th1-driven immune responses. Previous SAR investigations in several scaffolds of small molecule TLR7/8 activators pointed to the strict dependence of the selectivity for TLR7 vis-à-vis TLR8 on the electronic configurations of the heterocyclic systems, which we sought to examine quantitatively with the goal of developing "heuristics" to define structural requisites governing activity at TLR7 and/or TLR8. We undertook a scaffold-hopping approach, entailing the syntheses and biological evaluations of 13 different chemotypes. Crystal structures of TLR8 in complex with the two most active compounds confirmed important binding interactions playing a key role in ligand occupancy and biological activity. Density functional theory based quantum chemical calculations on these compounds followed by linear discriminant analyses permitted the classification of inactive, TLR8-active, and TLR7/8 dual-active compounds, confirming the critical role of partial charges in determining biological activity.
Organizational Affiliation:
Department of Medicinal Chemistry, University of Kansas , Multidisciplinary Research Building, Room 320D, 2030 Becker Drive, Lawrence, Kansas 66047, United States.