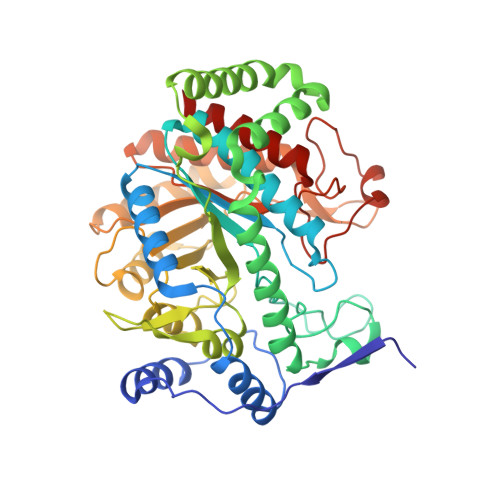
Probing the Sophisticated Synergistic Allosteric Regulation of Aromatic Amino Acid Biosynthesis in Mycobacterium tuberculosis Using -Amino Acids.
Reichau, S., Blackmore, N.J., Jiao, W., Parker, E.J.(2016) PLoS One 11: e0152723-e0152723
- PubMed: 27128682
- DOI: https://doi.org/10.1371/journal.pone.0152723
- Primary Citation of Related Structures:
5E2L, 5E40, 5E4N, 5E5G, 5E7Z - PubMed Abstract:
Chirality plays a major role in recognition and interaction of biologically important molecules. The enzyme 3-deoxy-d-arabino-heptulosonate 7-phosphate synthase (DAH7PS) is the first enzyme of the shikimate pathway, which is responsible for the synthesis of aromatic amino acids in bacteria and plants, and a potential target for the development of antibiotics and herbicides. DAH7PS from Mycobacterium tuberculosis (MtuDAH7PS) displays an unprecedented complexity of allosteric regulation, with three interdependent allosteric binding sites and a ternary allosteric response to combinations of the aromatic amino acids l-Trp, l-Phe and l-Tyr. In order to further investigate the intricacies of this system and identify key residues in the allosteric network of MtuDAH7PS, we studied the interaction of MtuDAH7PS with aromatic amino acids that bear the non-natural d-configuration, and showed that the d-amino acids do not elicit an allosteric response. We investigated the binding mode of d-amino acids using X-ray crystallography, site directed mutagenesis and isothermal titration calorimetry. Key differences in the binding mode were identified: in the Phe site, a hydrogen bond between the amino group of the allosteric ligands to the side chain of Asn175 is not established due to the inverted configuration of the ligands. In the Trp site, d-Trp forms no interaction with the main chain carbonyl group of Thr240 and less favourable interactions with Asn237 when compared to the l-Trp binding mode. Investigation of the MtuDAH7PSN175A variant further supports the hypothesis that the lack of key interactions in the binding mode of the aromatic d-amino acids are responsible for the absence of an allosteric response, which gives further insight into which residues of MtuDAH7PS play a key role in the transduction of the allosteric signal.
Organizational Affiliation:
Biomolecular Interaction Centre and Department of Chemistry, University of Canterbury, Christchurch, New Zealand.