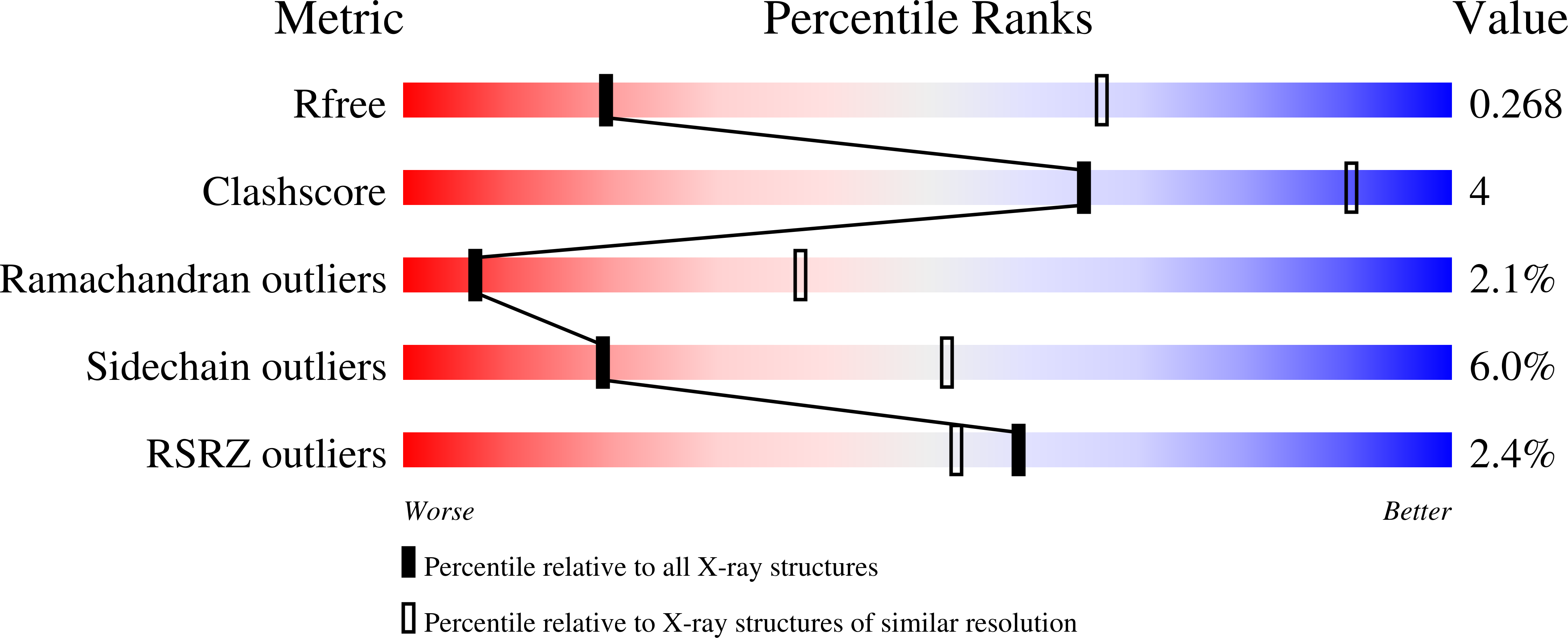
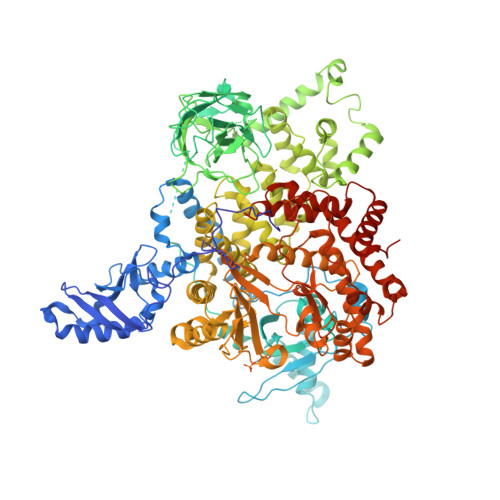
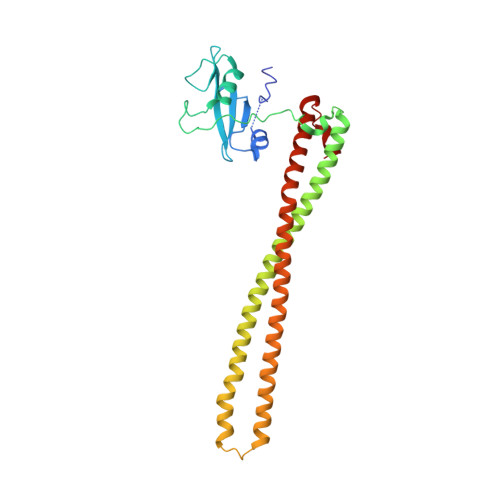
Identification of allosteric binding sites for PI3K alpha oncogenic mutant specific inhibitor design.
Miller, M.S., Maheshwari, S., McRobb, F.M., Kinzler, K.W., Amzel, L.M., Vogelstein, B., Gabelli, S.B.(2017) Bioorg Med Chem 25: 1481-1486
- PubMed: 28129991
- DOI: https://doi.org/10.1016/j.bmc.2017.01.012
- Primary Citation of Related Structures:
5SW8, 5SWG, 5SWO, 5SWP, 5SWR, 5SWT, 5SX8, 5SX9, 5SXA, 5SXB, 5SXC, 5SXD, 5SXE, 5SXF, 5SXI, 5SXJ, 5SXK - PubMed Abstract:
PIK3CA, the gene that encodes the catalytic subunit of phosphatidylinositol 3-kinase α (PI3Kα), is frequently mutated in breast and other types of cancer. A specific inhibitor that targets the mutant forms of PI3Kα could maximize treatment efficiency while minimizing side-effects. Herein we describe the identification of novel binding pockets that may provide an opportunity for the design of mutant selective inhibitors. Using a fragment-based approach, we screened a library of 352 fragments (MW<300Da) for binding to PI3Kα by X-ray crystallography. Five novel binding pockets were identified, each providing potential opportunities for inhibitor design. Of particular interest was a binding pocket near Glu542, which is located in one of the two most frequently mutated domains.
Organizational Affiliation:
Department of Oncology, Johns Hopkins University School of Medicine, Baltimore, MD 21205, United States.