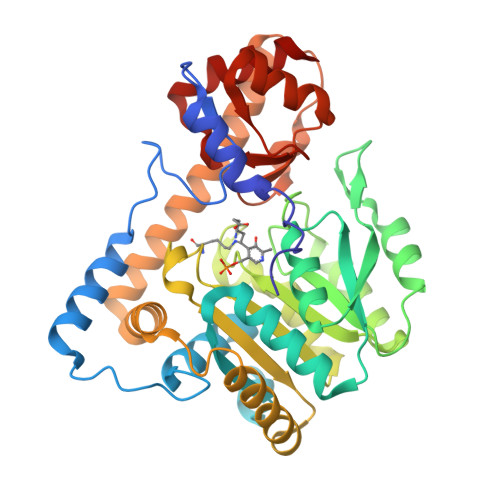
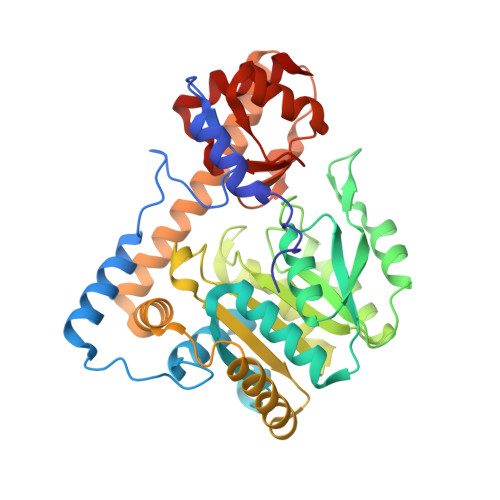
PLP and GABA trigger GabR-mediated transcription regulation in Bacillus subtilis via external aldimine formation.
Wu, R., Sanishvili, R., Belitsky, B.R., Juncosa, J.I., Le, H.V., Lehrer, H.J., Farley, M., Silverman, R.B., Petsko, G.A., Ringe, D., Liu, D.(2017) Proc Natl Acad Sci U S A 114: 3891-3896
- PubMed: 28348215
- DOI: https://doi.org/10.1073/pnas.1703019114
- Primary Citation of Related Structures:
5T4J, 5T4K, 5T4L - PubMed Abstract:
The Bacillus subtilis protein regulator of the gabTD operon and its own gene (GabR) is a transcriptional activator that regulates transcription of γ-aminobutyric acid aminotransferase (GABA-AT; GabT) upon interactions with pyridoxal-5'-phosphate (PLP) and GABA, and thereby promotes the biosynthesis of glutamate from GABA. We show here that the external aldimine formed between PLP and GABA is apparently responsible for triggering the GabR-mediated transcription activation. Details of the "active site" in the structure of the GabR effector-binding/oligomerization (Eb/O) domain suggest that binding a monocarboxylic γ-amino acid such as GABA should be preferred over dicarboxylic acid ligands. A reactive GABA analog, ( S )-4-amino-5-fluoropentanoic acid (AFPA), was used as a molecular probe to examine the reactivity of PLP in both GabR and a homologous aspartate aminotransferase (Asp-AT) from Escherichia coli as a control. A comparison between the structures of the Eb/O-PLP-AFPA complex and Asp-AT-PLP-AFPA complex revealed that GabR is incapable of facilitating further steps of the transamination reaction after the formation of the external aldimine. Results of in vitro and in vivo assays using full-length GabR support the conclusion that AFPA is an agonistic ligand capable of triggering GabR-mediated transcription activation via formation of an external aldimine with PLP.
Organizational Affiliation:
Department of Chemistry and Biochemistry, Loyola University Chicago, Chicago, IL 60660.