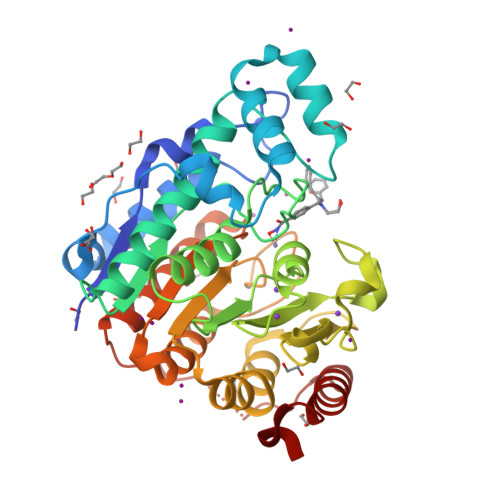
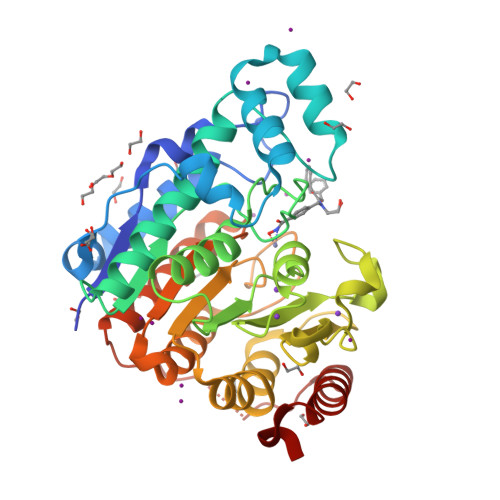
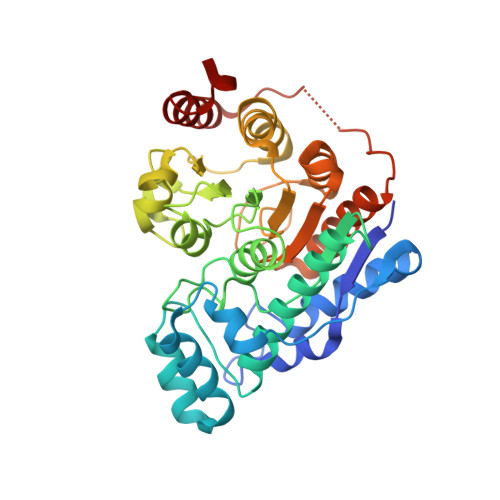
Unusual zinc-binding mode of HDAC6-selective hydroxamate inhibitors.
Porter, N.J., Mahendran, A., Breslow, R., Christianson, D.W.(2017) Proc Natl Acad Sci U S A 114: 13459-13464
- PubMed: 29203661
- DOI: https://doi.org/10.1073/pnas.1718823114
- Primary Citation of Related Structures:
5WGI, 5WGK, 5WGL, 5WGM - PubMed Abstract:
Histone deacetylases (HDACs) regulate myriad cellular processes by catalyzing the hydrolysis of acetyl-l-lysine residues in histone and nonhistone proteins. The Zn 2+ -dependent class IIb enzyme HDAC6 regulates microtubule function by deacetylating α-tubulin, which suppresses microtubule dynamics and leads to cell cycle arrest and apoptosis. Accordingly, HDAC6 is a target for the development of selective inhibitors that might be useful in new therapeutic approaches for the treatment of cancer, neurodegenerative diseases, and other disorders. Here, we present high-resolution structures of catalytic domain 2 from Danio rerio HDAC6 (henceforth simply "HDAC6") complexed with compounds that selectively inhibit HDAC6 while maintaining nanomolar inhibitory potency: N -hydroxy-4-[( N (2-hydroxyethyl)-2-phenylacetamido)methyl)-benzamide)] (HPB), ACY-1215 (Ricolinostat), and ACY-1083. These structures reveal that an unusual monodentate Zn 2+ coordination mode is exploited by sterically bulky HDAC6-selective phenylhydroxamate inhibitors. We additionally report the ultrahigh-resolution structure of the HDAC6-trichostatin A complex, which reveals two Zn 2+ -binding conformers for the inhibitor: a major conformer (70%) with canonical bidentate hydroxamate-Zn 2+ coordination geometry and a minor conformer (30%) with monodentate hydroxamate-Zn 2+ coordination geometry, reflecting a free energy difference of only 0.5 kcal/mol. The minor conformer is not visible in lower resolution structure determinations. Structural comparisons of HDAC6-inhibitor complexes with class I HDACs suggest active site features that contribute to the isozyme selectivity observed in biochemical assays.
Organizational Affiliation:
Roy and Diana Vagelos Laboratories, Department of Chemistry, University of Pennsylvania, Philadelphia, PA 19104-6323.