Discovery and Optimization of Inhibitors of the Parkinson's Disease Associated Protein DJ-1.
Tashiro, S., Caaveiro, J.M.M., Nakakido, M., Tanabe, A., Nagatoishi, S., Tamura, Y., Matsuda, N., Liu, D., Hoang, Q.Q., Tsumoto, K.(2018) ACS Chem Biol 13: 2783-2793
- PubMed: 30063823
- DOI: https://doi.org/10.1021/acschembio.8b00701
- Primary Citation of Related Structures:
6AF5, 6AF7, 6AF9, 6AFA, 6AFB, 6AFC, 6AFD, 6AFE, 6AFF, 6AFG, 6AFH, 6AFI, 6AFJ, 6AFL - PubMed Abstract:
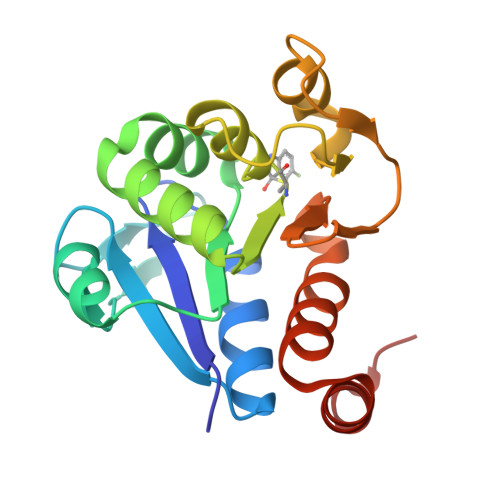
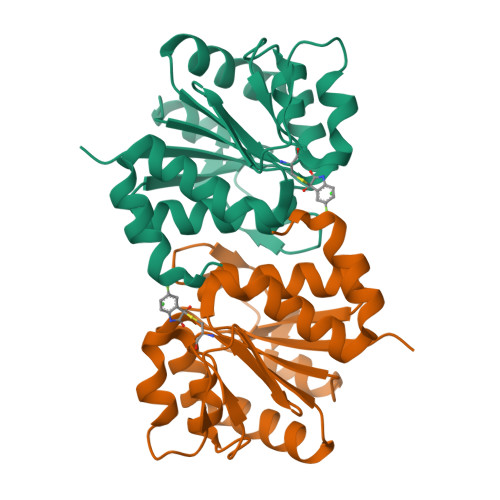
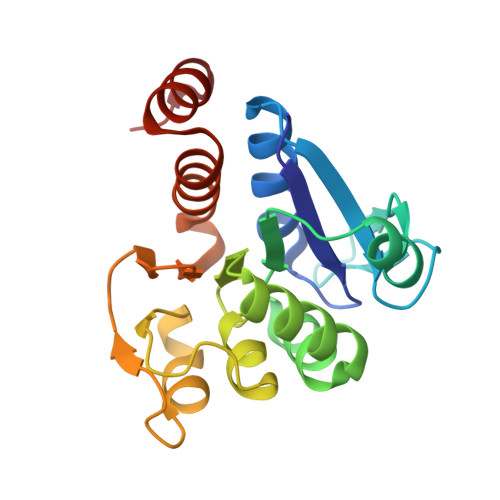
DJ-1 is a Parkinson's disease associated protein endowed with enzymatic, redox sensing, regulatory, chaperoning, and neuroprotective activities. Although DJ-1 has been vigorously studied for the past decade and a half, its exact role in the progression of the disease remains uncertain. In addition, little is known about the spatiotemporal regulation of DJ-1, or the biochemical basis explaining its numerous biological functions. Progress has been hampered by the lack of inhibitors with precisely known mechanisms of action. Herein, we have employed biophysical methodologies and X-ray crystallography to identify and to optimize a family of compounds inactivating the critical Cys106 residue of human DJ-1. We demonstrate these compounds are potent inhibitors of various activities of DJ-1 in vitro and in cell-based assays. This study reports a new family of DJ-1 inhibitors with a defined mechanism of action, and contributes toward the understanding of the biological function of DJ-1.
Organizational Affiliation:
Department of Bioengineering, Graduate School of Engineering , The University of Tokyo , Tokyo 113-8656 , Japan.