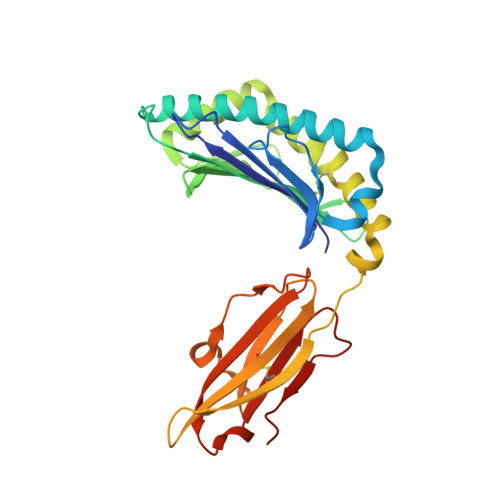
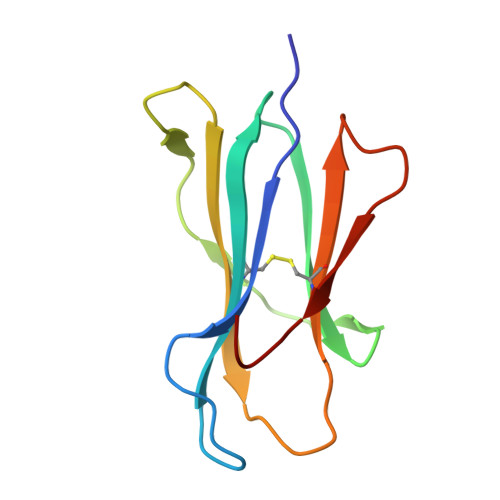
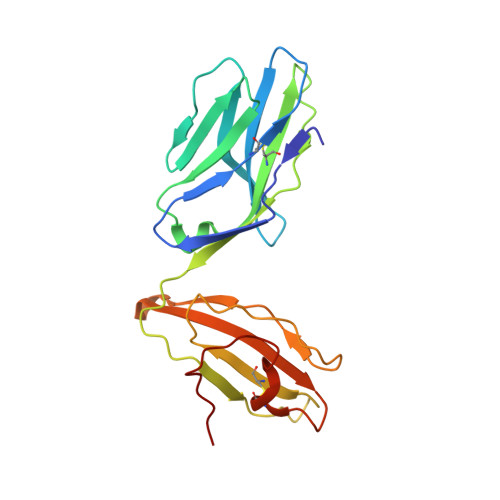
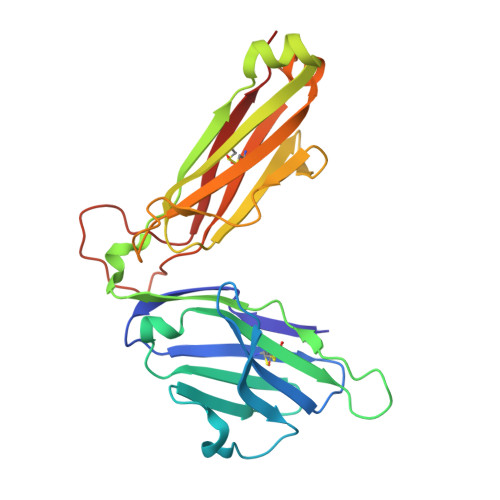
Isolation of a Structural Mechanism for Uncoupling T Cell Receptor Signaling from Peptide-MHC Binding.
Sibener, L.V., Fernandes, R.A., Kolawole, E.M., Carbone, C.B., Liu, F., McAffee, D., Birnbaum, M.E., Yang, X., Su, L.F., Yu, W., Dong, S., Gee, M.H., Jude, K.M., Davis, M.M., Groves, J.T., Goddard III, W.A., Heath, J.R., Evavold, B.D., Vale, R.D., Garcia, K.C.(2018) Cell 174: 672-687.e27
- PubMed: 30053426
- DOI: https://doi.org/10.1016/j.cell.2018.06.017
- Primary Citation of Related Structures:
6BJ2, 6BJ3, 6BJ8 - PubMed Abstract:
TCR-signaling strength generally correlates with peptide-MHC binding affinity; however, exceptions exist. We find high-affinity, yet non-stimulatory, interactions occur with high frequency in the human T cell repertoire. Here, we studied human TCRs that are refractory to activation by pMHC ligands despite robust binding. Analysis of 3D affinity, 2D dwell time, and crystal structures of stimulatory versus non-stimulatory TCR-pMHC interactions failed to account for their different signaling outcomes. Using yeast pMHC display, we identified peptide agonists of a formerly non-responsive TCR. Single-molecule force measurements demonstrated the emergence of catch bonds in the activating TCR-pMHC interactions, correlating with exclusion of CD45 from the TCR-APC contact site. Molecular dynamics simulations of TCR-pMHC disengagement distinguished agonist from non-agonist ligands based on the acquisition of catch bonds within the TCR-pMHC interface. The isolation of catch bonds as a parameter mediating the coupling of TCR binding and signaling has important implications for TCR and antigen engineering for immunotherapy.
Organizational Affiliation:
Departments of Molecular and Cellular Physiology and Structural Biology, Stanford University School of Medicine, Stanford, CA 94305, USA; Immunology Graduate Program, Stanford University School of Medicine, Stanford, CA 94305, USA.