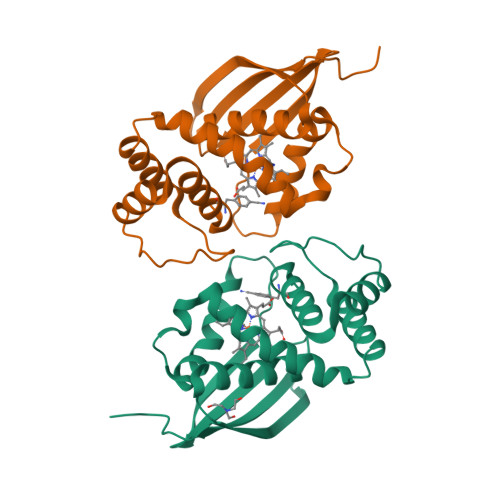
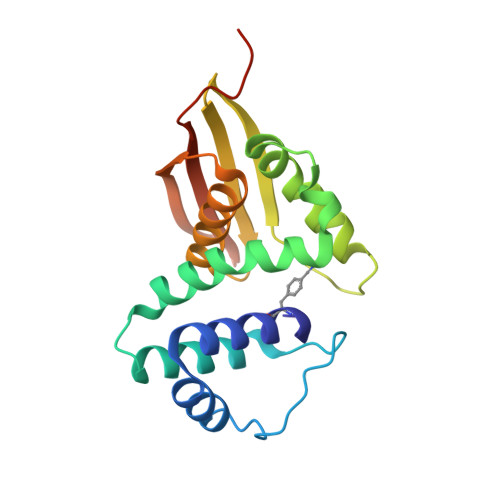
Exploring local solvation environments of a heme protein using the spectroscopic reporter 4-cyano-l-phenylalanine.
Kearney, C., Olenginski, L.T., Hirn, T.D., Fowler, G.D., Tariq, D., Brewer, S.H., Phillips-Piro, C.M.(2018) RSC Adv 8: 13503-13512
- PubMed: 29780583
- DOI: https://doi.org/10.1039/c8ra02000k
- Primary Citation of Related Structures:
6CWW - PubMed Abstract:
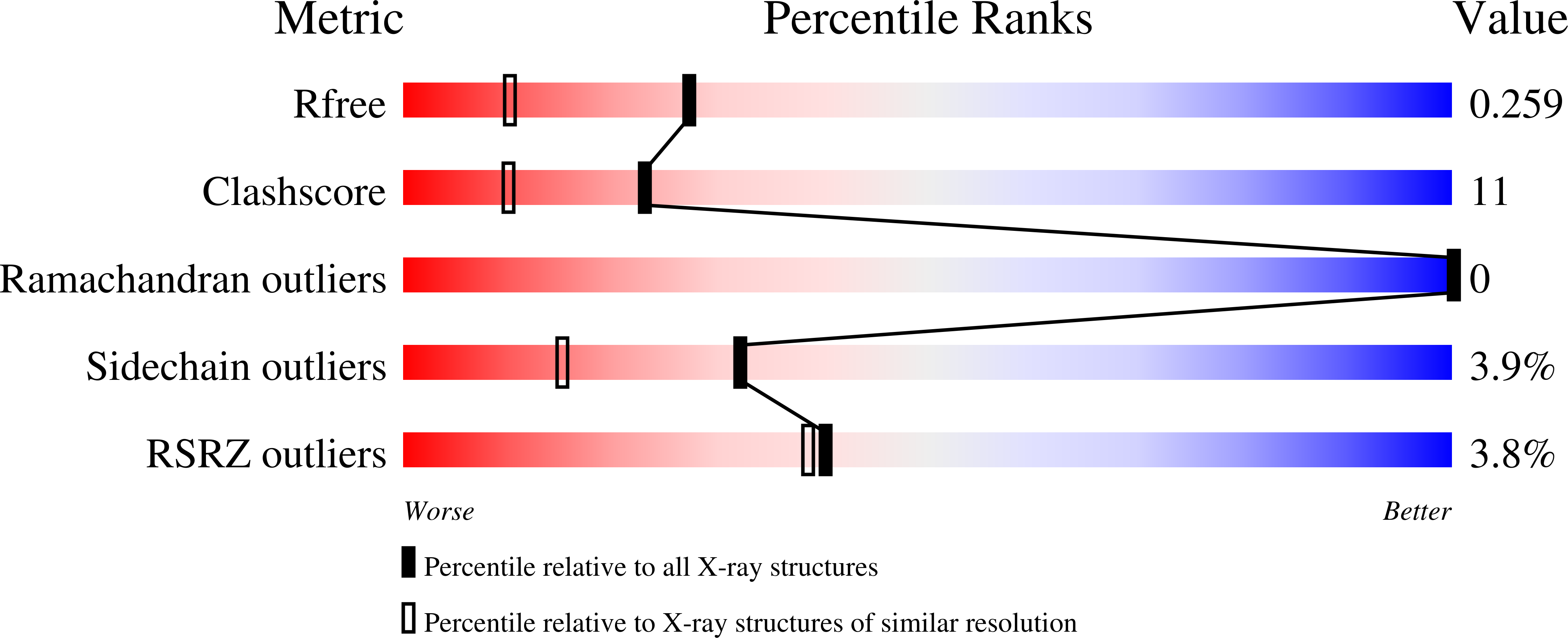
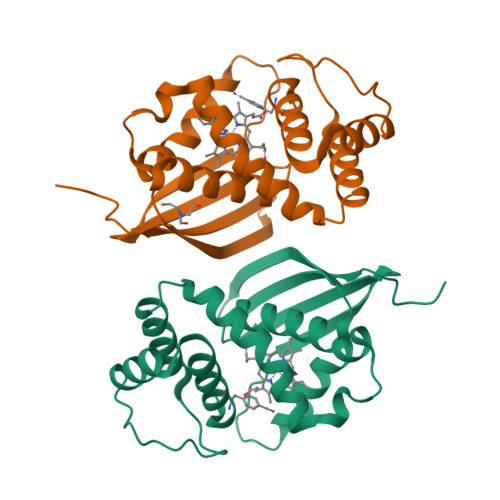
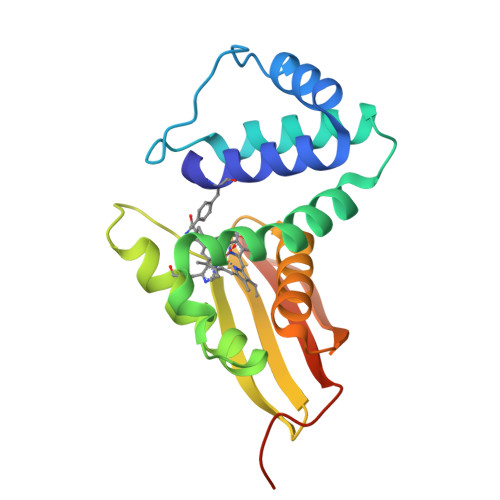
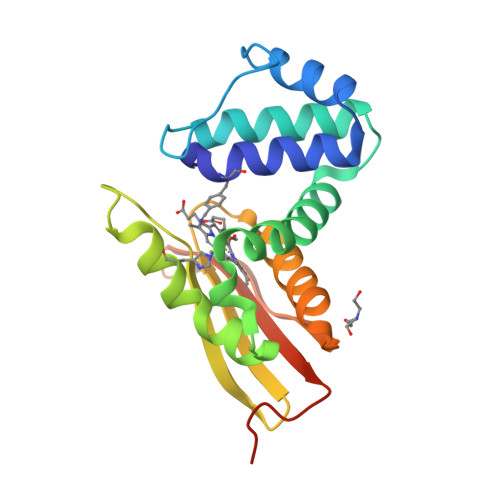
The vibrational reporter unnatural amino acid (UAA) 4-cyano-l-phenylalanine (pCNF) was genetically incorporated individually at three sites (5, 36, and 78) in the heme protein Caldanaerobacter subterraneus H-NOX to probe local hydration environments. The UAA pCNF was incorporated site-specifically using an engineered, orthogonal tRNA synthetase in E. coli . The ability of all of the pCNF-containing H-NOX proteins to form the ferrous CO, NO, or O 2 ligated and unligated states was confirmed with UV-Vis spectroscopy. The solvation state at each site of the three sites of pCNF incorporation was assessed using temperature-dependent infrared spectroscopy. Specifically, the frequency-temperature line slope (FTLS) method was utilized to show that the nitrile group at site 36 was fully solvated and the nitrile group at site 78 was de-solvated (buried) in the heme pocket. The nitrile group at site 5 was found to be partially solvated suggesting that the nitrile group was involved in moderate strength hydrogen bonds. These results were confirmed by the determination of the X-ray crystal structure of the H-NOX protein construct containing pCNF at site 5.
Organizational Affiliation:
Department of Chemistry, Franklin & Marshall College, PO Box 3003, Lancaster, PA 17604-3003, USA. Email: sbrewer@fandm.edu; Email: cpiro@fandm.edu.