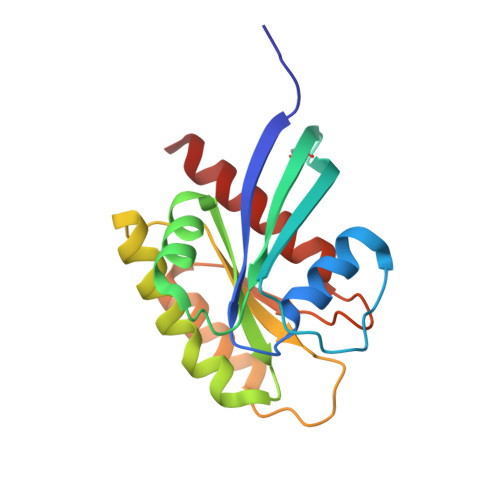
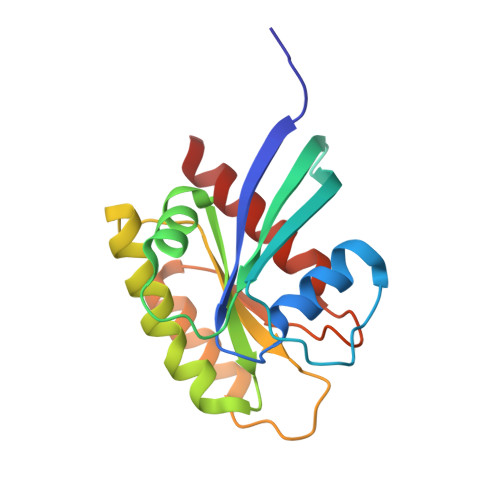
Small molecule inhibitors of RAS-effector protein interactions derived using an intracellular antibody fragment.
Quevedo, C.E., Cruz-Migoni, A., Bery, N., Miller, A., Tanaka, T., Petch, D., Bataille, C.J.R., Lee, L.Y.W., Fallon, P.S., Tulmin, H., Ehebauer, M.T., Fernandez-Fuentes, N., Russell, A.J., Carr, S.B., Phillips, S.E.V., Rabbitts, T.H.(2018) Nat Commun 9: 3169-3169
- PubMed: 30093669
- DOI: https://doi.org/10.1038/s41467-018-05707-2
- Primary Citation of Related Structures:
5OCG, 5OCO, 5OCT, 6FA1, 6FA2, 6FA3, 6FA4 - PubMed Abstract:
Targeting specific protein-protein interactions (PPIs) is an attractive concept for drug development, but hard to implement since intracellular antibodies do not penetrate cells and most small-molecule drugs are considered unsuitable for PPI inhibition. A potential solution to these problems is to select intracellular antibody fragments to block PPIs, use these antibody fragments for target validation in disease models and finally derive small molecules overlapping the antibody-binding site. Here, we explore this strategy using an anti-mutant RAS antibody fragment as a competitor in a small-molecule library screen for identifying RAS-binding compounds. The initial hits are optimized by structure-based design, resulting in potent RAS-binding compounds that interact with RAS inside the cells, prevent RAS-effector interactions and inhibit endogenous RAS-dependent signalling. Our results may aid RAS-dependent cancer drug development and demonstrate a general concept for developing small compounds to replace intracellular antibody fragments, enabling rational drug development to target validated PPIs.
Organizational Affiliation:
Weatherall Institute of Molecular Medicine, MRC Molecular Haematology Unit, University of Oxford, John Radcliffe Hospital, Oxford, OX3 9DS, UK.