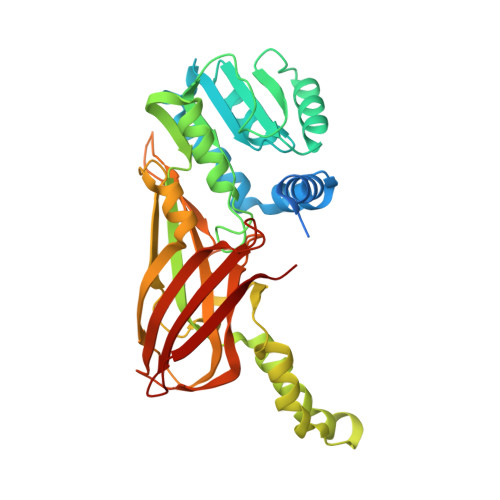
Discovery of a First-in-Class Protein Arginine Methyltransferase 6 (PRMT6) Covalent Inhibitor
Shen, Y., Li, F., Szewczyk, M.M., Halabelian, L., Park, K.S., Chau, I., Dong, A., Zeng, H., Chen, H., Meng, F., Barsyte-Lovejoy, D., Arrowsmith, C.H., Brown, P.J., Liu, J., Vedadi, M., Jin, J.(2020) J Med Chem 63: 5477-5487