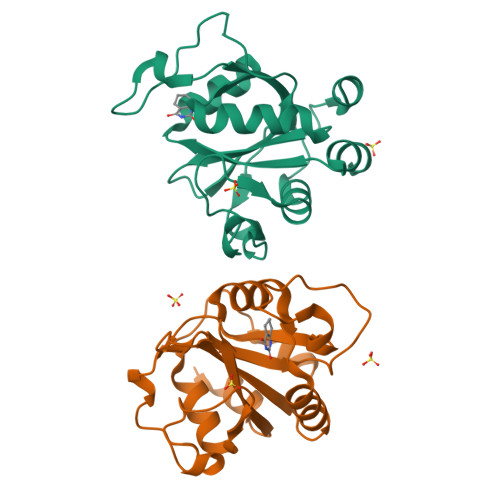
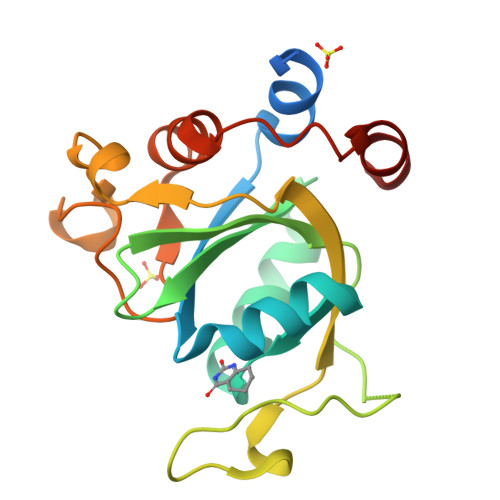
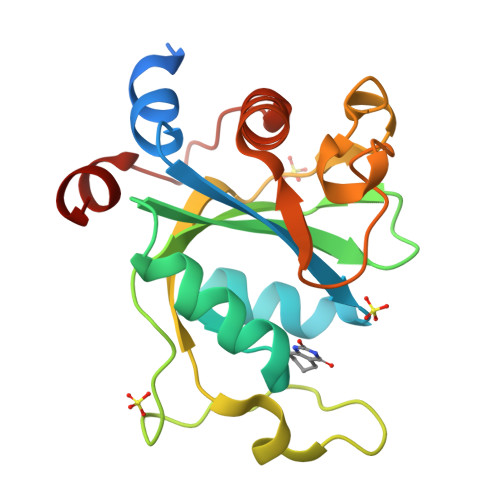
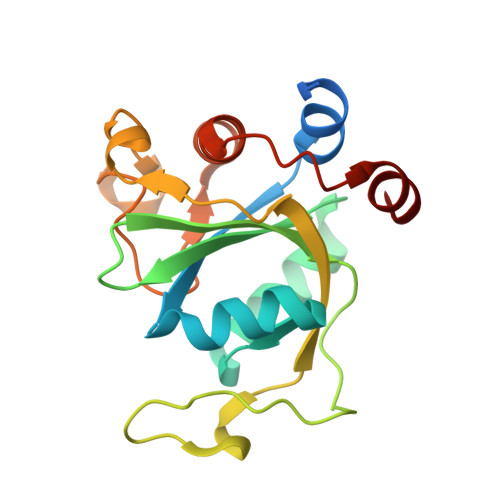
Selectively Disrupting m6A-Dependent Protein-RNA Interactions with Fragments.
Bedi, R.K., Huang, D., Wiedmer, L., Li, Y., Dolbois, A., Wojdyla, J.A., Sharpe, M.E., Caflisch, A., Sledz, P.(2020) ACS Chem Biol 15: 618-625
- PubMed: 32101404
- DOI: https://doi.org/10.1021/acschembio.9b00894
- Primary Citation of Related Structures:
6SYZ, 6SZ1, 6SZ2, 6SZ3, 6SZ7, 6SZ8, 6SZL, 6SZN, 6SZR, 6SZT, 6SZX, 6SZY, 6T01, 6T02, 6T03, 6T04, 6T05, 6T06, 6T07, 6T08, 6T09, 6T0A, 6T0C, 6T0D, 6T0O, 6T0X, 6T0Z, 6T10, 6T11, 6T12 - PubMed Abstract:
We report a crystallographic analysis of small-molecule ligands of the human YTHDC1 domain that recognizes N6-methylated adenine (m 6 A) in RNA. The 30 binders are fragments (molecular weight < 300 g mol -1 ) that represent 10 different chemotypes identified by virtual screening. Despite the structural disorder of the binding site loop (residues 429-439), most of the 30 fragments emulate the two main interactions of the -NHCH 3 group of m 6 A. These interactions are the hydrogen bond to the backbone carbonyl of Ser378 and the van der Waals contacts with the tryptophan cage. Different chemical groups are involved in the conserved binding motifs. Some of the fragments show favorable ligand efficiency for YTHDC1 and selectivity against other m 6 A reader domains. The structural information is useful for the design of modulators of m 6 A recognition by YTHDC1.
Organizational Affiliation:
Department of Biochemistry, University of Zurich, Winterthurerstrasse 190, CH-8057 Zurich, Switzerland.