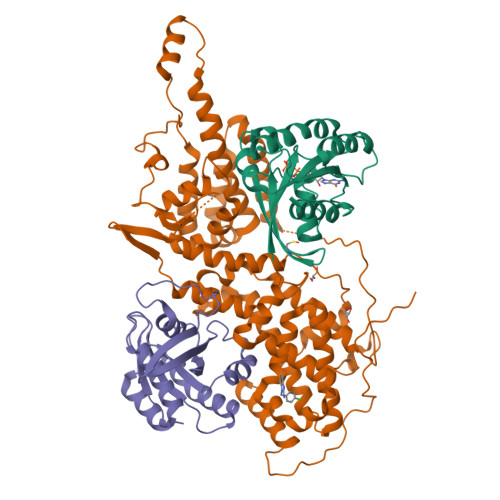
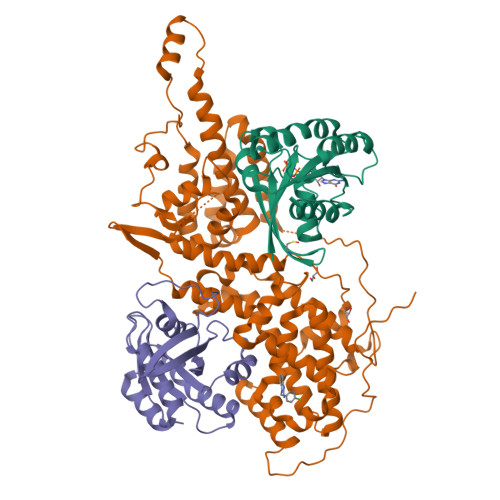
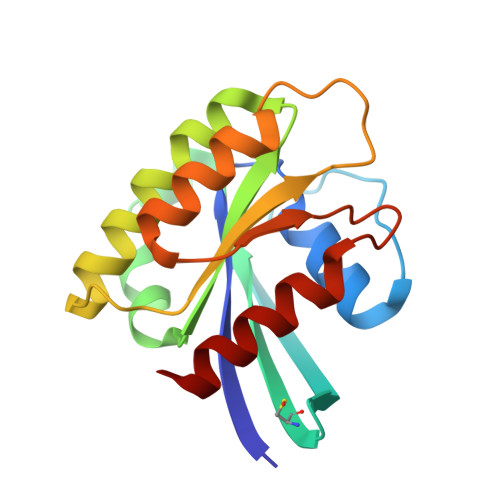
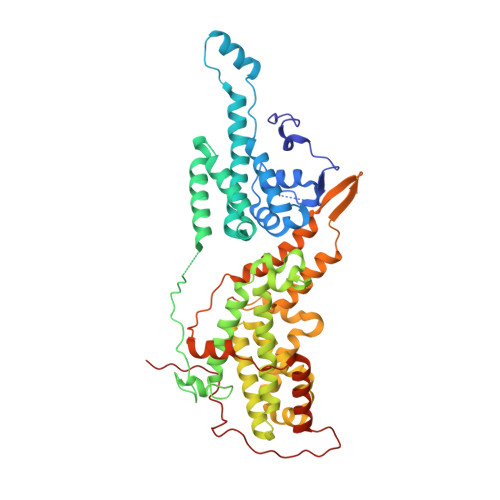
Discovery of Sulfonamide-Derived Agonists of SOS1-Mediated Nucleotide Exchange on RAS Using Fragment-Based Methods.
Sarkar, D., Olejniczak, E.T., Phan, J., Coker, J.A., Sai, J., Arnold, A., Beesetty, Y., Waterson, A.G., Fesik, S.W.(2020) J Med Chem 63: 8325-8337
- PubMed: 32673492
- DOI: https://doi.org/10.1021/acs.jmedchem.0c00511
- Primary Citation of Related Structures:
6V94, 6V9F, 6V9J, 6V9L, 6V9M, 6V9N - PubMed Abstract:
The nucleotide exchange factor Son of Sevenless (SOS) catalyzes the activation of RAS by converting it from its inactive GDP-bound state to its active GTP-bound state. Recently, we have reported the discovery of small-molecule allosteric activators of SOS1 that can increase the amount of RAS-GTP in cells. The compounds can inhibit ERK phosphorylation at higher concentrations by engaging a feedback mechanism. To further study this process, we sought different chemical matter from an NMR-based fragment screen using selective methyl labeling. To aid this process, several Ile methyl groups located in different binding sites of the protein were assigned and used to categorize the NMR hits into different classes. Hit to lead optimization using an iterative structure-based design paradigm resulted in compounds with improvements in binding affinity. These improved molecules of a different chemical class increase SOS1 cat -mediated nucleotide exchange on RAS and display cellular action consistent with our prior results.
Organizational Affiliation:
Department of Biochemistry, Vanderbilt University School of Medicine, Nashville, Tennessee 37232-0146, United States.