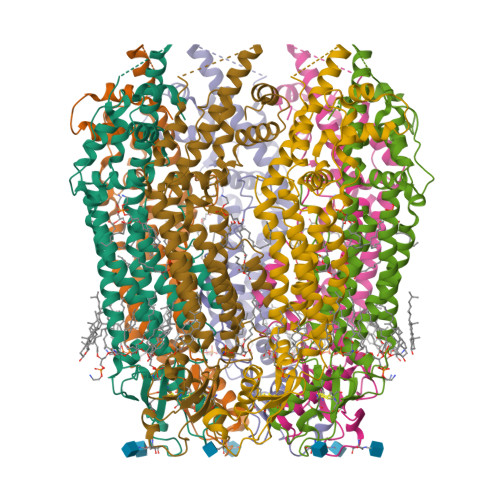
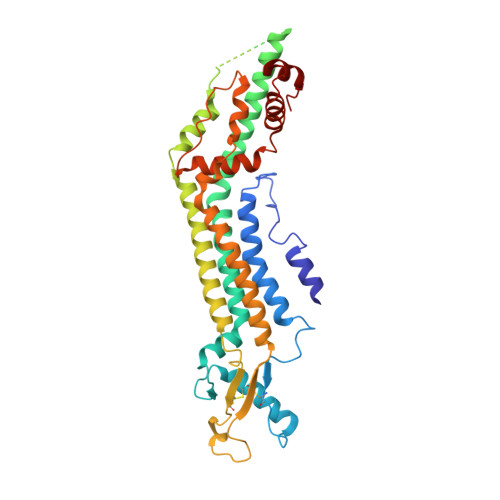
Structures of human pannexin 1 reveal ion pathways and mechanism of gating.
Ruan, Z., Orozco, I.J., Du, J., Lu, W.(2020) Nature 584: 646-651
- PubMed: 32494015
- DOI: https://doi.org/10.1038/s41586-020-2357-y
- Primary Citation of Related Structures:
6WBF, 6WBG, 6WBI, 6WBK, 6WBL, 6WBM, 6WBN - PubMed Abstract:
Pannexin 1 (PANX1) is an ATP-permeable channel with critical roles in a variety of physiological functions such as blood pressure regulation 1 , apoptotic cell clearance 2 and human oocyte development 3 . Here we present several structures of human PANX1 in a heptameric assembly at resolutions of up to 2.8 angström, including an apo state, a caspase-7-cleaved state and a carbenoxolone-bound state. We reveal a gating mechanism that involves two ion-conducting pathways. Under normal cellular conditions, the intracellular entry of the wide main pore is physically plugged by the C-terminal tail. Small anions are conducted through narrow tunnels in the intracellular domain. These tunnels connect to the main pore and are gated by a long linker between the N-terminal helix and the first transmembrane helix. During apoptosis, the C-terminal tail is cleaved by caspase, allowing the release of ATP through the main pore. We identified a carbenoxolone-binding site embraced by W74 in the extracellular entrance and a role for carbenoxolone as a channel blocker. We identified a gap-junction-like structure using a glycosylation-deficient mutant, N255A. Our studies provide a solid foundation for understanding the molecular mechanisms underlying the channel gating and inhibition of PANX1 and related large-pore channels.
Organizational Affiliation:
Van Andel Institute, Grand Rapids, MI, USA.