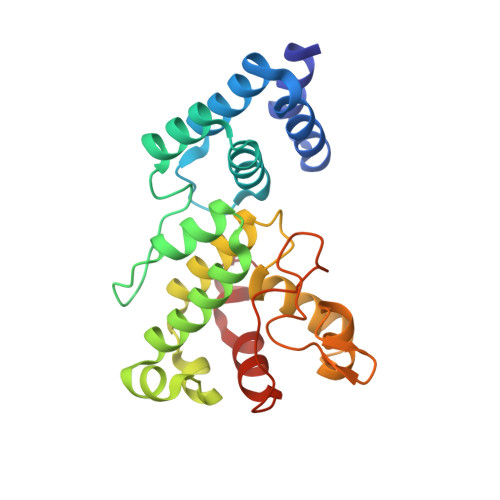
Structural basis of bilin binding by the chlorophyll biosynthesis regulator GUN4.
Hu, J.H., Chang, J.W., Xu, T., Wang, J., Wang, X., Lin, R., Duanmu, D., Liu, L.(2021) Protein Sci 30: 2083-2091
- PubMed: 34382282
- DOI: https://doi.org/10.1002/pro.4164
- Primary Citation of Related Structures:
7E2R, 7E2S, 7E2T, 7E2U - PubMed Abstract:
The chlorophyll biosynthesis regulator GENOMES UNCOUPLED 4 (GUN4) is conserved in nearly all oxygenic photosynthetic organisms. Recently, GUN4 has been found to be able to bind the linear tetrapyrroles (bilins) and stimulate the magnesium chelatase activity in the unicellular green alga Chlamydomonas reinhardtii. Here, we characterize GUN4 proteins from Arabidopsis thaliana and the cyanobacterium Synechocystis sp. PCC 6803 for their ability to bind bilins, and present the crystal structures of Synechocystis GUN4 in biliverdin-bound, phycocyanobilin-bound, and phytochromobilin-bound forms at the resolutions of 1.05, 1.10, and 1.70 Å, respectively. These linear molecules adopt a cyclic-helical conformation, and bind more tightly than planar porphyrins to the tetrapyrrole-binding pocket of GUN4. Based on structural comparison, we propose a working model of GUN4 in regulation of tetrapyrrole biosynthetic pathway, and address the role of the bilin-bound GUN4 in retrograde signaling.
Organizational Affiliation:
School of Life Sciences, Anhui University, Hefei, China.