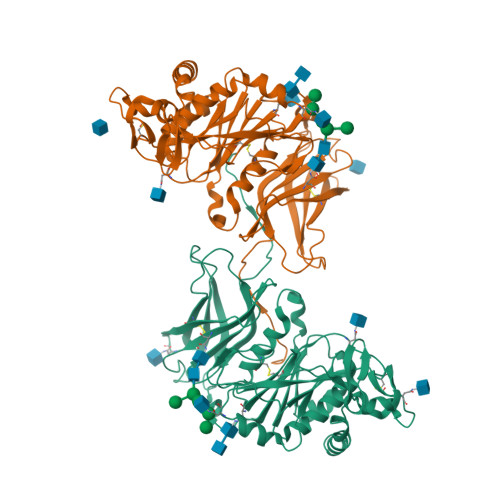
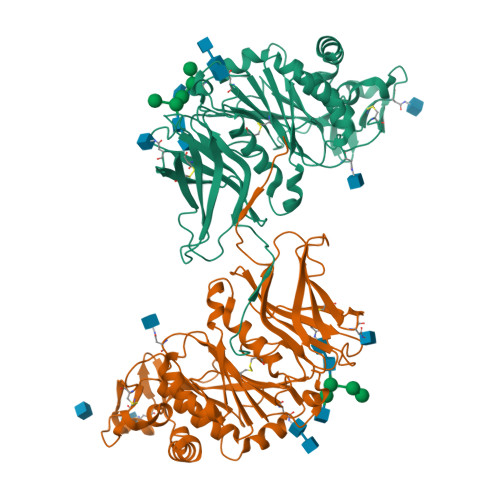
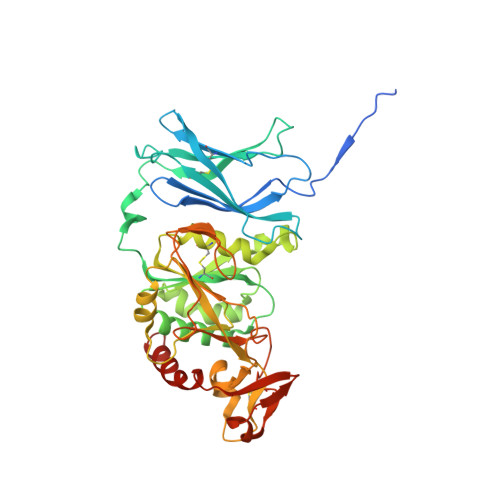
Structural and biochemical insight into a modular beta-1,4-galactan synthase in plants.
Prabhakar, P.K., Pereira, J.H., Taujale, R., Shao, W., Bharadwaj, V.S., Chapla, D., Yang, J.Y., Bomble, Y.J., Moremen, K.W., Kannan, N., Hammel, M., Adams, P.D., Scheller, H.V., Urbanowicz, B.R.(2023) Nat Plants 9: 486-500
- PubMed: 36849618
- DOI: https://doi.org/10.1038/s41477-023-01358-4
- Primary Citation of Related Structures:
8D3T, 8D3Z - PubMed Abstract:
Rhamnogalacturonan I (RGI) is a structurally complex pectic polysaccharide with a backbone of alternating rhamnose and galacturonic acid residues substituted with arabinan and galactan side chains. Galactan synthase 1 (GalS1) transfers galactose and arabinose to either extend or cap the β-1,4-galactan side chains of RGI, respectively. Here we report the structure of GalS1 from Populus trichocarpa, showing a modular protein consisting of an N-terminal domain that represents the founding member of a new family of carbohydrate-binding module, CBM95, and a C-terminal glycosyltransferase family 92 (GT92) catalytic domain that adopts a GT-A fold. GalS1 exists as a dimer in vitro, with stem domains interacting across the chains in a 'handshake' orientation that is essential for maintaining stability and activity. In addition to understanding the enzymatic mechanism of GalS1, we gained insight into the donor and acceptor substrate binding sites using deep evolutionary analysis, molecular simulations and biochemical studies. Combining all the results, a mechanism for GalS1 catalysis and a new model for pectic galactan side-chain addition are proposed.
Organizational Affiliation:
Department of Biochemistry and Molecular Biology, University of Georgia, Athens, GA, USA.