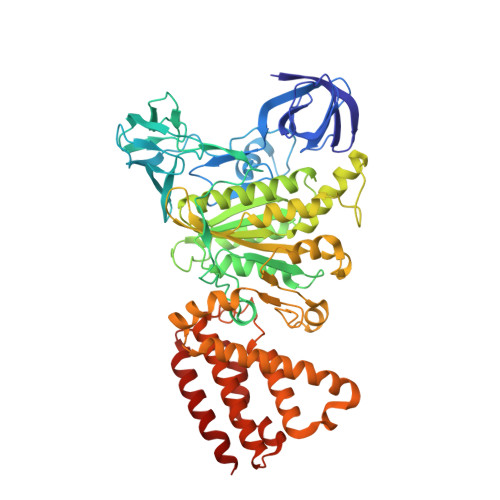
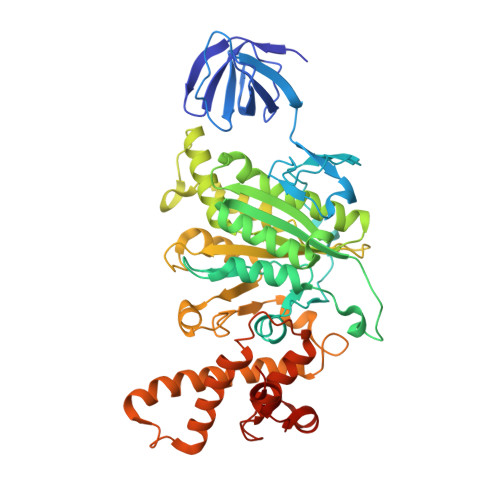
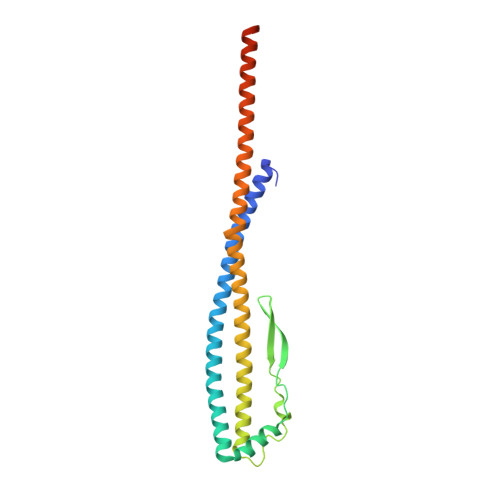
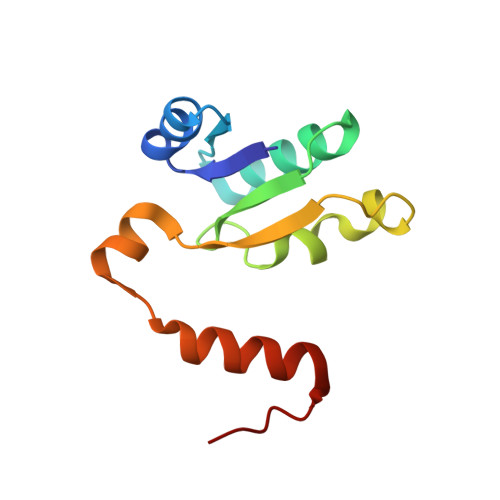
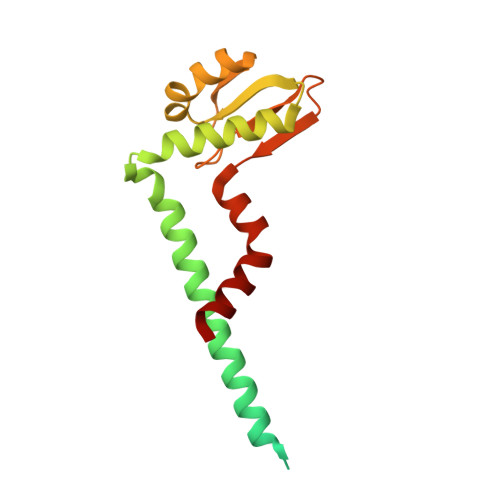
Cryo-EM analysis of V/A-ATPase intermediates reveals the transition of the ground-state structure to steady-state structures by sequential ATP binding.
Nakanishi, A., Kishikawa, J.I., Mitsuoka, K., Yokoyama, K.(2023) J Biol Chem 299: 102884-102884
- PubMed: 36626983
- DOI: https://doi.org/10.1016/j.jbc.2023.102884
- Primary Citation of Related Structures:
8GXU, 8GXW, 8GXX, 8GXY, 8GXZ - PubMed Abstract:
Vacuolar/archaeal-type ATPase (V/A-ATPase) is a rotary ATPase that shares a common rotary catalytic mechanism with F o F 1 ATP synthase. Structural images of V/A-ATPase obtained by single-particle cryo-electron microscopy during ATP hydrolysis identified several intermediates, revealing the rotary mechanism under steady-state conditions. However, further characterization is needed to understand the transition from the ground state to the steady state. Here, we identified the cryo-electron microscopy structures of V/A-ATPase corresponding to short-lived initial intermediates during the activation of the ground state structure by time-resolving snapshot analysis. These intermediate structures provide insights into how the ground-state structure changes to the active, steady state through the sequential binding of ATP to its three catalytic sites. All the intermediate structures of V/A-ATPase adopt the same asymmetric structure, whereas the three catalytic dimers adopt different conformations. This is significantly different from the initial activation process of F o F 1 , where the overall structure of the F 1 domain changes during the transition from a pseudo-symmetric to a canonical asymmetric structure (PNAS NEXUS, pgac116, 2022). In conclusion, our findings provide dynamical information that will enhance the future prospects for studying the initial activation processes of the enzymes, which have unknown intermediate structures in their functional pathway.
Organizational Affiliation:
Department of Molecular Biosciences, Kyoto Sangyo University, Kamigamo-Motoyama, Kita-ku, Kyoto, Japan; Research Center for Ultra-High Voltage Electron Microscopy, Osaka University, Osaka, Japan.