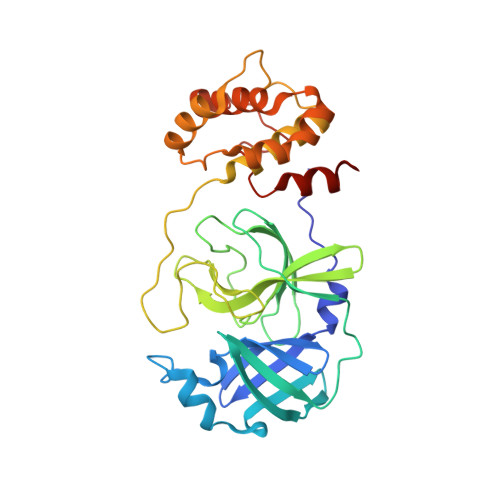
The H163A mutation unravels an oxidized conformation of the SARS-CoV-2 main protease.
Tran, N., Dasari, S., Barwell, S.A.E., McLeod, M.J., Kalyaanamoorthy, S., Holyoak, T., Ganesan, A.(2023) Nat Commun 14: 5625-5625
- PubMed: 37699927
- DOI: https://doi.org/10.1038/s41467-023-40023-4
- Primary Citation of Related Structures:
8DD6, 8DDL, 8SG6 - PubMed Abstract:
The main protease of SARS-CoV-2 (Mpro) is an important target for developing COVID-19 therapeutics. Recent work has highlighted Mpro's susceptibility to undergo redox-associated conformational changes in response to cellular and immune-system-induced oxidation. Despite structural evidence indicating large-scale rearrangements upon oxidation, the mechanisms of conformational change and its functional consequences are poorly understood. Here, we present the crystal structure of an Mpro point mutant (H163A) that shows an oxidized conformation with the catalytic cysteine in a disulfide bond. We hypothesize that Mpro adopts this conformation under oxidative stress to protect against over-oxidation. Our metadynamics simulations illustrate a potential mechanism by which H163 modulates this transition and suggest that this equilibrium exists in the wild type enzyme. We show that other point mutations also significantly shift the equilibrium towards this state by altering conformational free energies. Unique avenues of SARS-CoV-2 research can be explored by understanding how H163 modulates this equilibrium.
Organizational Affiliation:
Department of Biology, Faculty of Science, University of Waterloo, 200 University Avenue West, Waterloo, ON, N2L 3G1, Canada.