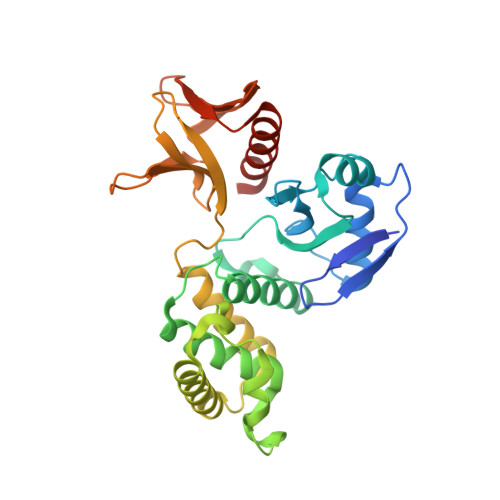
Structural analysis of the FERM domain of human protein tyrosine phosphatase non-receptor type 21.
Lee, H.S., Ku, B., Shin, H.C., Kim, S.J.(2024) Acta Crystallogr F Struct Biol Commun 80: 148-153
- PubMed: 38940939
- DOI: https://doi.org/10.1107/S2053230X24005260
- Primary Citation of Related Structures:
8Y8Y - PubMed Abstract:
Protein tyrosine phosphatase non-receptor type 21 (PTPN21) is a cytosolic protein tyrosine phosphatase that regulates cell growth and invasion. Due to its oncogenic properties, PTPN21 has recently emerged as a potential therapeutic target for cancer. In this study, the three-dimensional structure of the PTPN21 FERM domain was determined at 2.1 Å resolution by X-ray crystallography. The crystal structure showed that this domain harbors canonical FERM folding and consists of three subdomains that are tightly packed via highly conserved intramolecular hydrophobic interactions. Consistent with this, the PTPN21 FERM domain shares high structural homology with several other FERM domains. Moreover, structural superimposition demonstrated two putative protein-binding sites of the PTPN21 FERM domain, which are presumed to be associated with interaction with its binding partner, kinesin family member 1C. Thus, these data suggest that the FERM domain of PTPN21 serves as a module that mediates protein-protein interaction, like other FERM domains.
Organizational Affiliation:
Disease Target Structure Research Center, Korea Research Institute of Bioscience and Biotechnology, Daejeon 34141, Republic of Korea.