Potency-enhanced peptidomimetic VHL ligands with improved oral bioavailability
Wu, H., Murray, J.M., Ishisoko, N., Frommlet, A., Fairbrother, W.J., Fuhrmann, J.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Elongin-B | 104 | Homo sapiens | Mutation(s): 0 Gene Names: ELOB, TCEB2 | 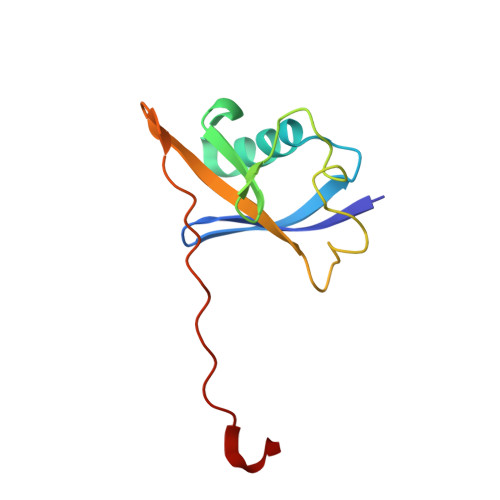 | |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for Q15370 (Homo sapiens) Explore Q15370 Go to UniProtKB: Q15370 | |||||
PHAROS: Q15370 GTEx: ENSG00000103363 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q15370 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Elongin-C | 96 | Homo sapiens | Mutation(s): 0 Gene Names: ELOC, TCEB1 | 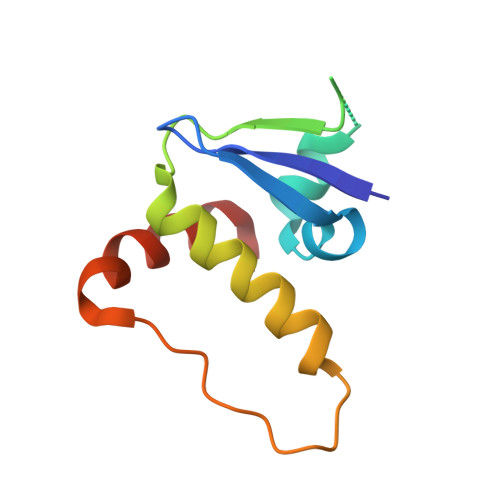 | |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for Q15369 (Homo sapiens) Explore Q15369 Go to UniProtKB: Q15369 | |||||
PHAROS: Q15369 GTEx: ENSG00000154582 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q15369 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| von Hippel-Lindau disease tumor suppressor | 176 | Homo sapiens | Mutation(s): 0 Gene Names: VHL | 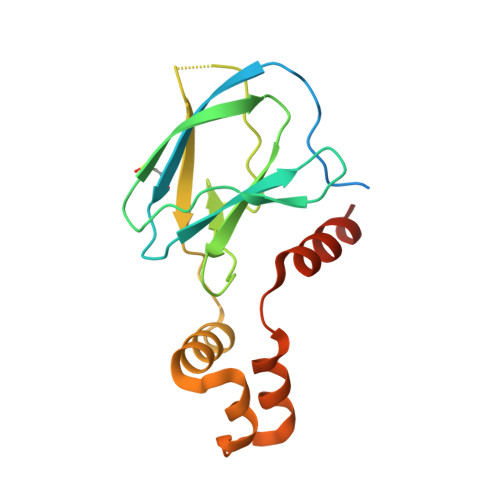 | |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for P40337 (Homo sapiens) Explore P40337 Go to UniProtKB: P40337 | |||||
PHAROS: P40337 GTEx: ENSG00000134086 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P40337 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 4 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| von Hippel-Lindau disease tumor suppressor | 176 | Homo sapiens | Mutation(s): 0 Gene Names: VHL | 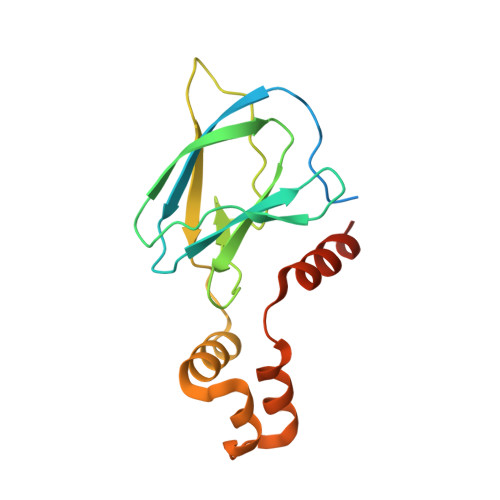 | |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for P40337 (Homo sapiens) Explore P40337 Go to UniProtKB: P40337 | |||||
PHAROS: P40337 GTEx: ENSG00000134086 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P40337 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Find similar proteins by: Sequence | 3D Structure
Entity ID: 5 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| 3-PYRIDIN-4-YL-2,4-DIHYDRO-INDENO[1,2-.C.]PYRAZOLE | M [auth W], N [auth X], O [auth Y], P [auth Z] | 5 | synthetic construct | Mutation(s): 0 | 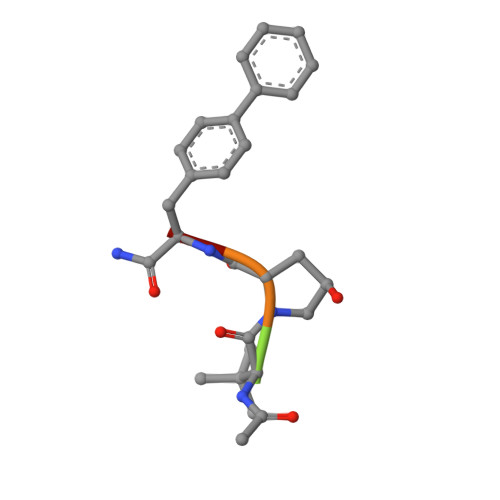 |
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 5 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| TFA Query on TFA | Q [auth B], Z [auth K] | trifluoroacetic acid C2 H F3 O2 DTQVDTLACAAQTR-UHFFFAOYSA-N |  | ||
| PEG Query on PEG | R [auth B] | DI(HYDROXYETHYL)ETHER C4 H10 O3 MTHSVFCYNBDYFN-UHFFFAOYSA-N |  | ||
| DMS Query on DMS | AA [auth L], W [auth H], X [auth I] | DIMETHYL SULFOXIDE C2 H6 O S IAZDPXIOMUYVGZ-UHFFFAOYSA-N |  | ||
| EDO Query on EDO | S [auth C], Y [auth K] | 1,2-ETHANEDIOL C2 H6 O2 LYCAIKOWRPUZTN-UHFFFAOYSA-N |  | ||
| ACT Query on ACT | T [auth C], U [auth F], V [auth H] | ACETATE ION C2 H3 O2 QTBSBXVTEAMEQO-UHFFFAOYSA-M |  | ||
| Modified Residues 3 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Type | Formula | 2D Diagram | Parent |
| CSX Query on CSX | C, L | L-PEPTIDE LINKING | C3 H7 N O3 S |  | CYS |
| HYP Query on HYP | M [auth W], N [auth X], O [auth Y], P [auth Z] | L-PEPTIDE LINKING | C5 H9 N O3 |  | PRO |
| TBG Query on TBG | M [auth W], N [auth X], O [auth Y], P [auth Z] | L-PEPTIDE LINKING | C6 H13 N O2 |  | VAL |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 93.297 | α = 90 |
| b = 93.297 | β = 90 |
| c = 362.757 | γ = 90 |
| Software Name | Purpose |
|---|---|
| REFMAC | refinement |
| HKL-2000 | data scaling |
| HKL-2000 | data reduction |
| PHASER | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Not funded | -- |