A novel series of positive modulators of the AMPA receptor: structure-based lead optimization.
Jamieson, C., Campbell, R.A., Cumming, I.A., Gillen, K.J., Gillespie, J., Kazemier, B., Kiczun, M., Lamont, Y., Lyons, A.J., Maclean, J.K., Martin, F., Moir, E.M., Morrow, J.A., Pantling, J., Rankovic, Z., Smith, L.(2010) Bioorg Med Chem Lett 20: 6072-6075
- PubMed: 20817521
- DOI: https://doi.org/10.1016/j.bmcl.2010.08.063
- Primary Citation of Related Structures:
3O6G, 3O6H, 3O6I - PubMed Abstract:
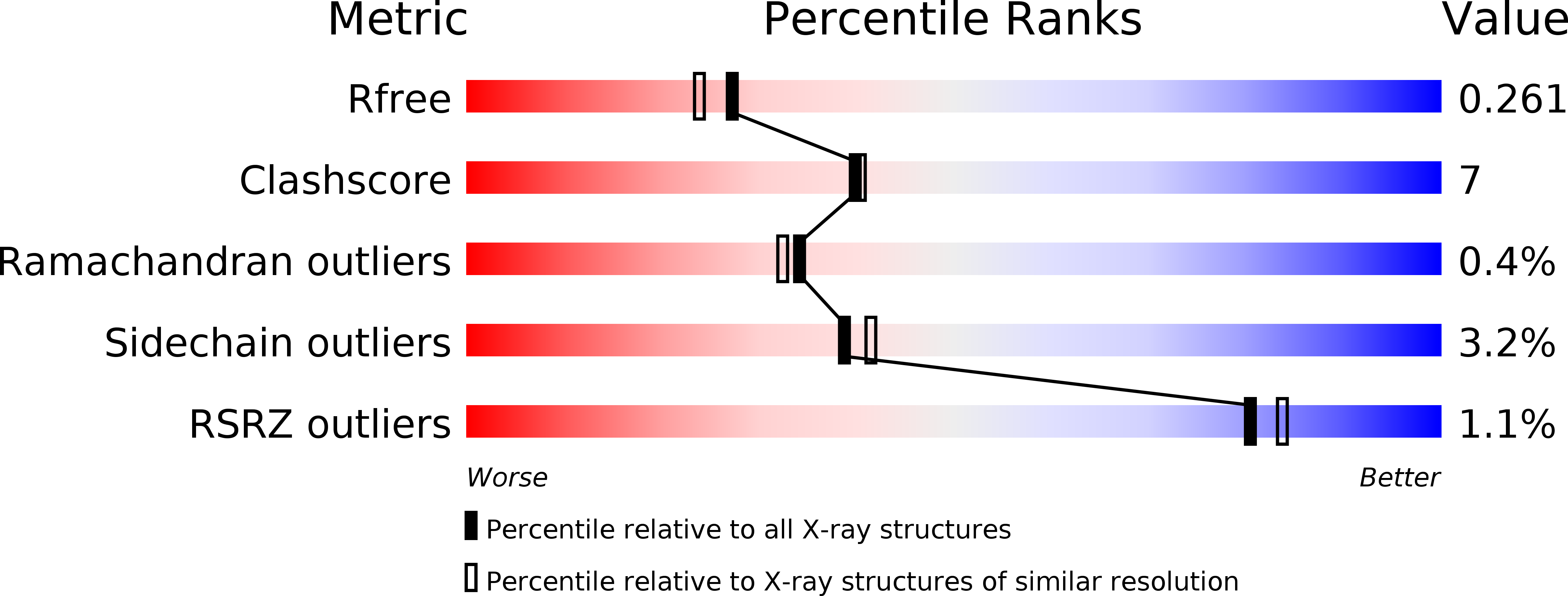
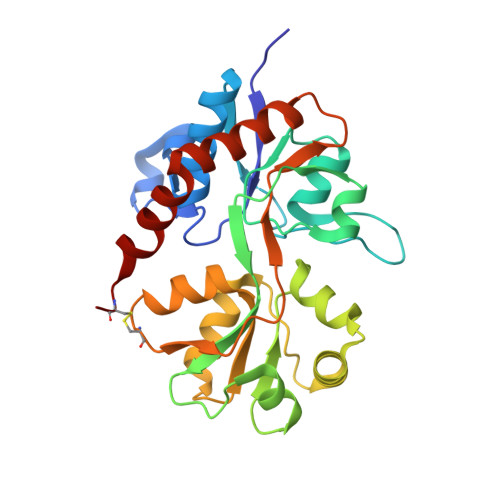
Starting from lead compound 1, we demonstrate how X-ray structural data can be used to understand SAR and expediently optimize bioavailability in a novel series of AMPA receptor modulators, furnishing 5 with improved bioavailability and robust in vivo activity.
Organizational Affiliation:
Merck Research Laboratories, MSD, Newhouse, Motherwell, Lanarkshire, UK. craig.jamieson@strath.ac.uk