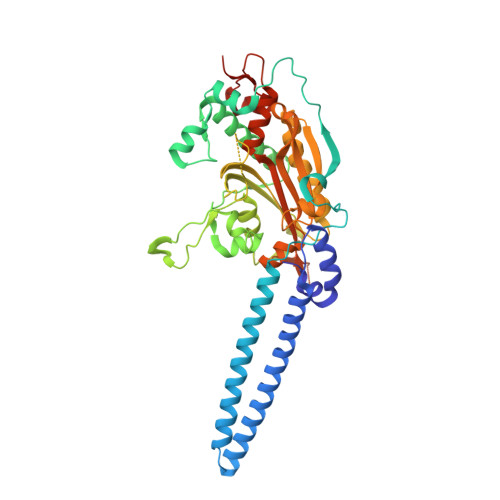
Structure-Guided Enhancement of Selectivity of Chemical Probe Inhibitors Targeting Bacterial Seryl-tRNA Synthetase.
Cain, R., Salimraj, R., Punekar, A.S., Bellini, D., Fishwick, C.W.G., Czaplewski, L., Scott, D.J., Harris, G., Dowson, C.G., Lloyd, A.J., Roper, D.I.(2019) J Med Chem 62: 9703-9717
- PubMed: 31626547
- DOI: https://doi.org/10.1021/acs.jmedchem.9b01131
- Primary Citation of Related Structures:
6R1M, 6R1N, 6R1O - PubMed Abstract:
Aminoacyl-tRNA synthetases are ubiquitous and essential enzymes for protein synthesis and also a variety of other metabolic processes, especially in bacterial species. Bacterial aminoacyl-tRNA synthetases represent attractive and validated targets for antimicrobial drug discovery if issues of prokaryotic versus eukaryotic selectivity and antibiotic resistance generation can be addressed. We have determined high-resolution X-ray crystal structures of the Escherichia coli and Staphylococcus aureus seryl-tRNA synthetases in complex with aminoacyl adenylate analogues and applied a structure-based drug discovery approach to explore and identify a series of small molecule inhibitors that selectively inhibit bacterial seryl-tRNA synthetases with greater than 2 orders of magnitude compared to their human homologue, demonstrating a route to the selective chemical inhibition of these bacterial targets.
Organizational Affiliation:
School of Life Sciences , University of Warwick , Gibbet Hill Road , Coventry CV4 7AL , United Kingdom.