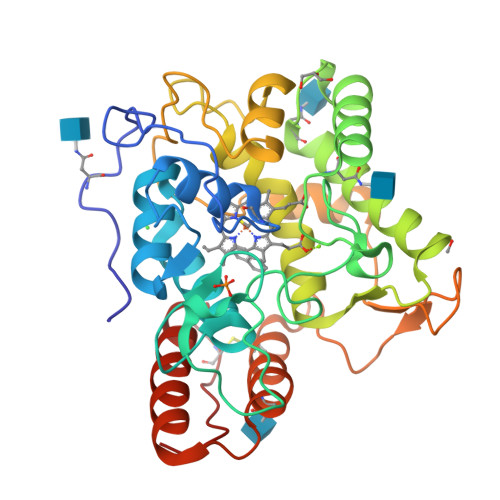
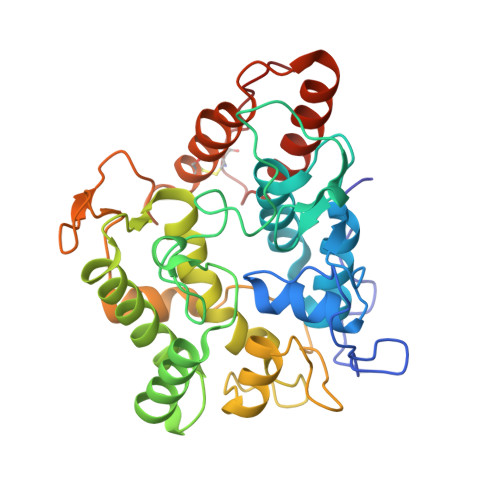
Engineering a Highly Regioselective Fungal Peroxygenase for the Synthesis of Hydroxy Fatty Acids.
Gomez de Santos, P., Gonzalez-Benjumea, A., Fernandez-Garcia, A., Aranda, C., Wu, Y., But, A., Molina-Espeja, P., Mate, D.M., Gonzalez-Perez, D., Zhang, W., Kiebist, J., Scheibner, K., Hofrichter, M., Swiderek, K., Moliner, V., Sanz-Aparicio, J., Hollmann, F., Gutierrez, A., Alcalde, M.(2023) Angew Chem Int Ed Engl 62: e202217372-e202217372
- PubMed: 36583658
- DOI: https://doi.org/10.1002/anie.202217372
- Primary Citation of Related Structures:
7PN4, 7PN5, 7PN6, 7PN7, 7PN8, 7PN9, 7PNA - PubMed Abstract:
The hydroxylation of fatty acids is an appealing reaction in synthetic chemistry, although the lack of selective catalysts hampers its industrial implementation. In this study, we have engineered a highly regioselective fungal peroxygenase for the ω-1 hydroxylation of fatty acids with quenched stepwise over-oxidation. One single mutation near the Phe catalytic tripod narrowed the heme cavity, promoting a dramatic shift toward subterminal hydroxylation with a drop in the over-oxidation activity. While crystallographic soaking experiments and molecular dynamic simulations shed light on this unique oxidation pattern, the selective biocatalyst was produced by Pichia pastoris at 0.4 g L -1 in a fed-batch bioreactor and used in the preparative synthesis of 1.4 g of (ω-1)-hydroxytetradecanoic acid with 95 % regioselectivity and 83 % ee for the S enantiomer.
Organizational Affiliation:
Evoenzyme, S.L., Parque Científico de Madrid, C/Faraday 7, 28049, Madrid, Spain.