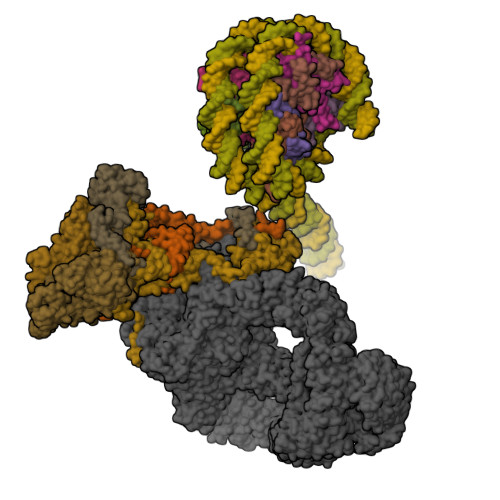
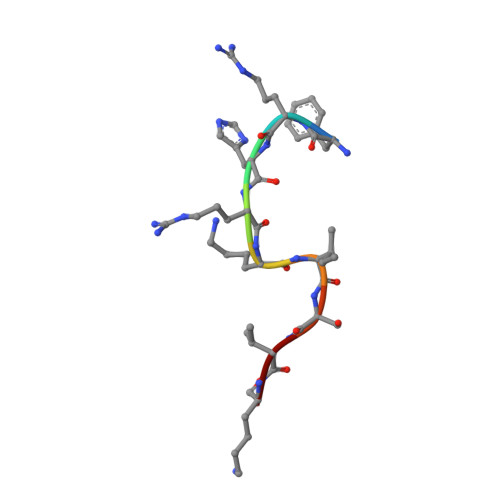
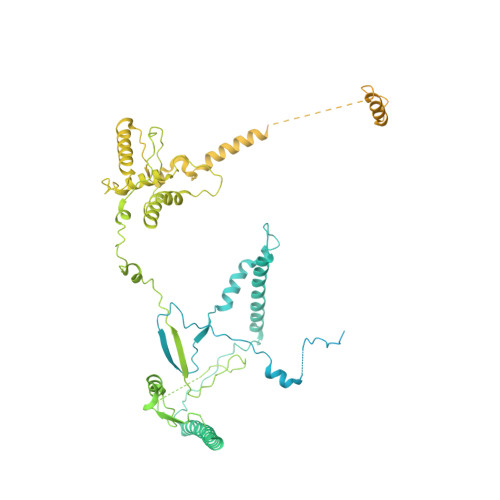
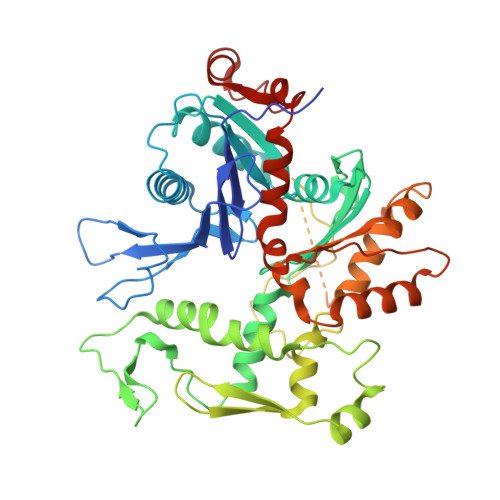
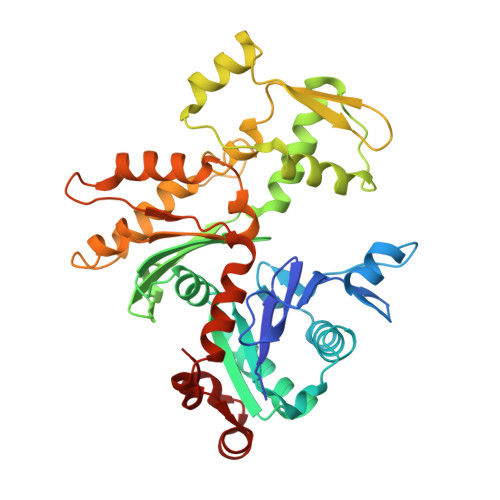
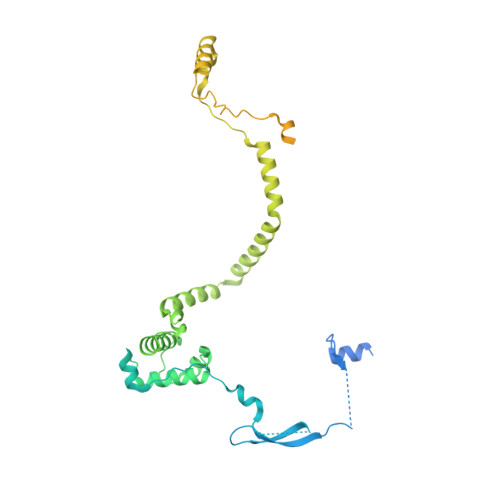
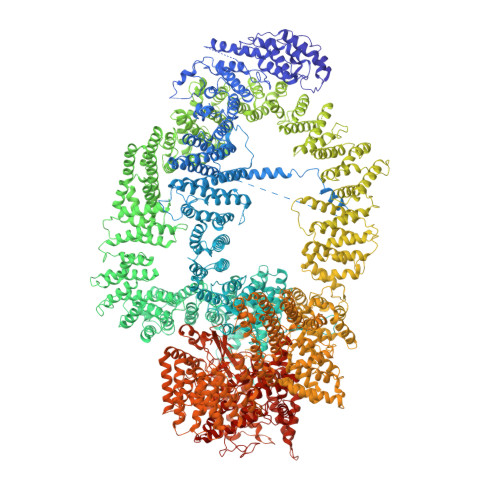
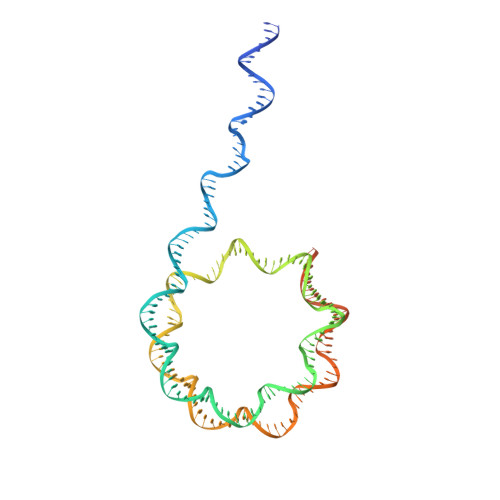
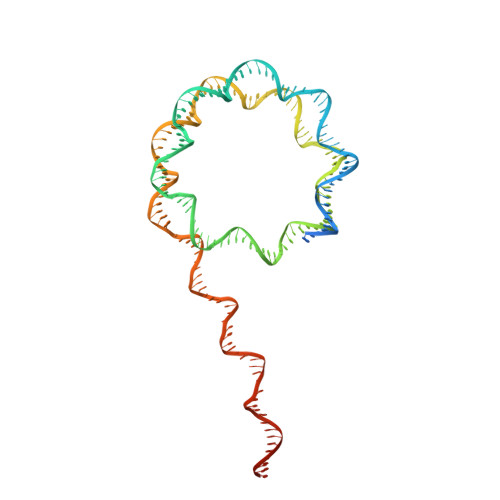
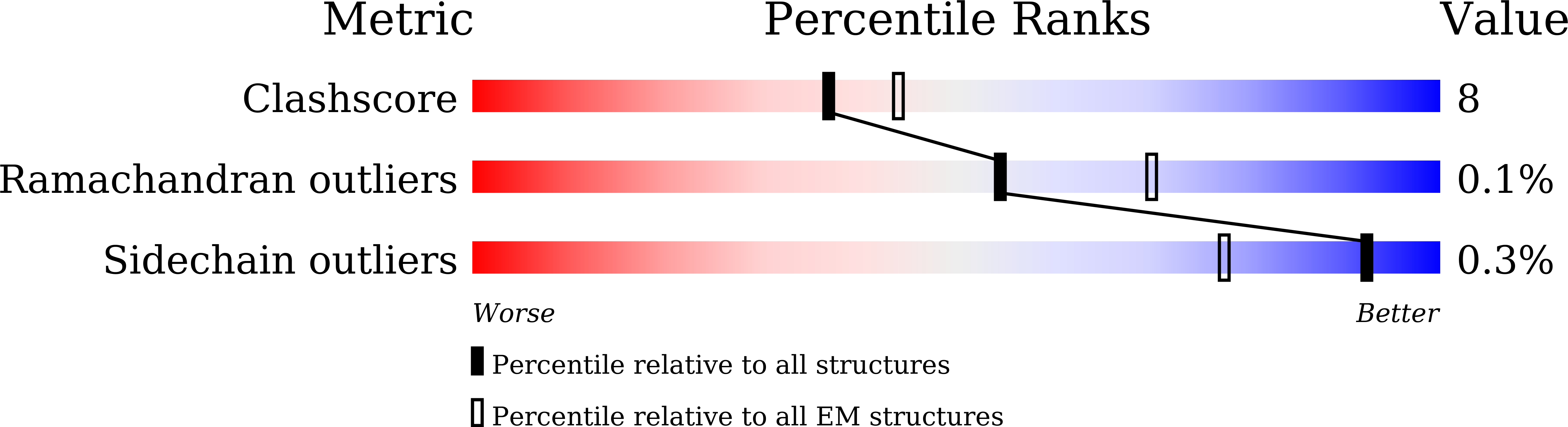Structure of the NuA4 acetyltransferase complex bound to the nucleosome.
Qu, K., Chen, K., Wang, H., Li, X., Chen, Z.(2022) Nature 610: 569-574
- PubMed: 36198799
- DOI: https://doi.org/10.1038/s41586-022-05303-x
- Primary Citation of Related Structures:
7VVU, 7VVY, 7VVZ - PubMed Abstract:
Deoxyribonucleic acid in eukaryotes wraps around the histone octamer to form nucleosomes 1 , the fundamental unit of chromatin. The N termini of histone H4 interact with nearby nucleosomes and play an important role in the formation of high-order chromatin structure and heterochromatin silencing 2-4 . NuA4 in yeast and its homologue Tip60 complex in mammalian cells are the key enzymes that catalyse H4 acetylation, which in turn regulates chromatin packaging and function in transcription activation and DNA repair 5-10 . Here we report the cryo-electron microscopy structure of NuA4 from Saccharomyces cerevisiae bound to the nucleosome. NuA4 comprises two major modules: the catalytic histone acetyltransferase (HAT) module and the transcription activator-binding (TRA) module. The nucleosome is mainly bound by the HAT module and is positioned close to a polybasic surface of the TRA module, which is important for the optimal activity of NuA4. The nucleosomal linker DNA carrying the upstream activation sequence is oriented towards the conserved, transcription activator-binding surface of the Tra1 subunit, which suggests a potential mechanism of NuA4 to act as a transcription co-activator. The HAT module recognizes the disk face of the nucleosome through the H2A-H2B acidic patch and nucleosomal DNA, projecting the catalytic pocket of Esa1 to the N-terminal tail of H4 and supporting its function in selective acetylation of H4. Together, our findings illustrate how NuA4 is assembled and provide mechanistic insights into nucleosome recognition and transcription co-activation by a HAT.
Organizational Affiliation:
MOE Key Laboratory of Protein Science, Tsinghua University, Beijing, P. R. China.