News
Visualize Structure Quality Metrics in 3D
03/12
wwPDB Validation Reports are available for every entry to provide an assessment of the quality of a structure and highlight specific concerns by considering the model coordinates, experimental data, and fit between the two.
Use RCSB PDB's viewer (NGL) to display information from this report in 3D.
 NGL view using the Color by Random by Coil Index option (PDB structure 2n2t)
NGL view using the Color by Random by Coil Index option (PDB structure 2n2t)Color By Random Coil Index: For NMR structures with available chemical shift data, the "NMR Random Coil Index" (RCI) scheme colors a structure according to the probability that the given residue is disordered ("random coil-like"). The color of each residue indicates whether the residue is classified as rigid (blue, RCI = 0.0) or flexible (red, RCI = 0.6). Residue coloring is based on the measured chemical shifts and on the primary sequence of the protein chain (for more on RCI, see Berjanskii 2005 and Berjanskii 2008). Residues without chemical shift data are displayed in gray.
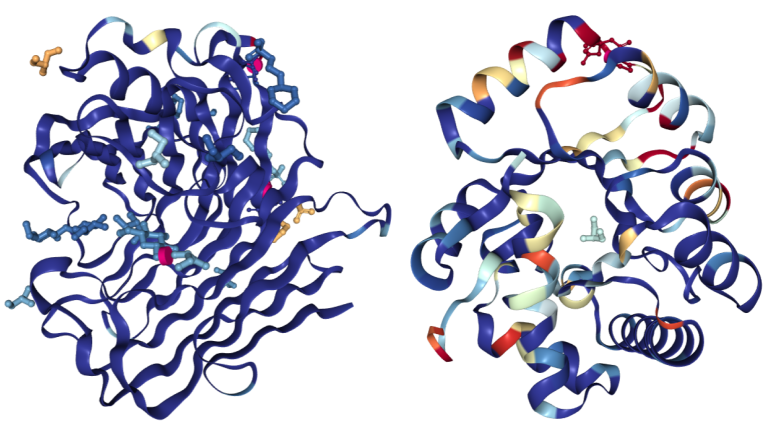 Hydrolases colored using the “Density Fit” scheme. On the left is a structure of Endoglucanase A (PDB structure 3WY6) with a generally good fit; on the right a structure of Ribonuclease P protein component 3 (PDB ID 3WYZ), which has areas of more problematic fit.
Hydrolases colored using the “Density Fit” scheme. On the left is a structure of Endoglucanase A (PDB structure 3WY6) with a generally good fit; on the right a structure of Ribonuclease P protein component 3 (PDB ID 3WYZ), which has areas of more problematic fit.Color By Density Fit: For structures determined using X-ray crystallography with available structure factors, “Density Fit” colors a structure according to the quality of agreement between the model and the experimental electron density. Blue indicates a good fit for a residue and red a poor fit. Residue coloring is determined using normalized Real Space R (RSRZ) for polymer residues and real space correlation coefficient (RSCC) for ligands. Colors range from red (RSRZ=-2 or RSCC=0.678), through white, to blue (RSRZ=0 or RSCC=1.0).
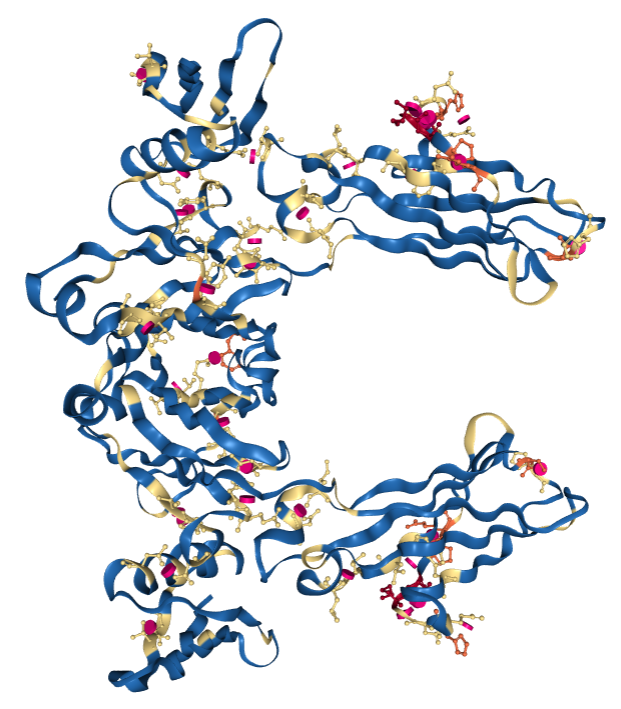 PDB structure1FCC, a 3.2Å resolution structure with worse overall quality relative to all X-ray structures colored by Geometry Quality.
PDB structure1FCC, a 3.2Å resolution structure with worse overall quality relative to all X-ray structures colored by Geometry Quality.Color By Geometry Quality: Color each polymer residue and ligand molecule according to the number of geometric issues (blue for 0, yellow for 1, orange for 2, and red for 3 or more). Protein residues and nucleotides are colored per residue whereas ligand molecules are colored per atom. Possible geometric issues include steric clashes, Ramachandran or RNA backbone outliers, and sidechain conformation outliers.
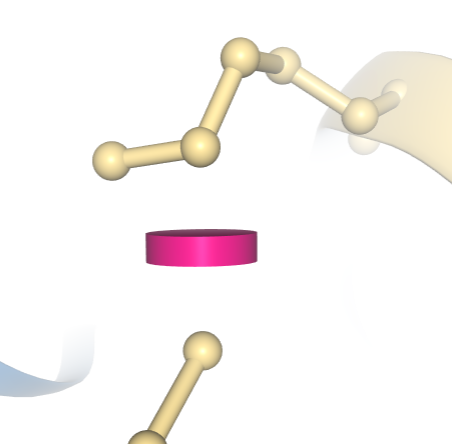 A clash between two atoms in PDB structure 1D66 is indicated by a pink disc, showing how much the atoms’ vdW spheres overlap.
A clash between two atoms in PDB structure 1D66 is indicated by a pink disc, showing how much the atoms’ vdW spheres overlap.Show/Hide Clashes: Select the Clashes checkbox to display clashes between pairs of atoms as pink discs, with the size of each disc reflecting the degree of van der Waals (vdW) overlap between the two atoms. Clash display is currently not available for structures comprising more than 10,000 residues.












